When omics data do not look perfectly normal, many researchers default to a nonparametric test. In practice, however, the bigger issue is often not normality alone, but unequal variance, unbalanced sample sizes, missing values, outliers, and the biological question you actually want to answer. This article explains what Student's t-test, Welch's t-test, and Mann-Whitney U each measure, why they are not interchangeable, and how to choose the right two-group test for proteomics, metabolomics, and other omics studies with more confidence.
1. In Real Omics Projects, the Problem Is Rarely Just Non-Normality
Searches such as "t test vs welch t test proteomics," "mann whitney u metabolomics," "two group test omics," "statistical test selection omics," and "omics data heteroscedasticity" all point to the same practical issue: in real omics datasets, the main challenge is rarely non-normality alone. More often, the real problem is unequal variance between groups, imbalanced sample sizes, missing-value handling, or a small number of biologically real yet statistically influential extreme observations.
A good example comes from serum metabolomics of pulmonary nodules. In a large Nature Communications study, the discovery cohort included healthy controls, benign nodules, and stage I lung adenocarcinoma, with pyruvate and lactate elevated in the cancer group; the authors used Wilcoxon rank tests for discovery screening. That choice is sensible when the goal is to detect distributional separation, but it does not answer exactly the same question as a mean-based analysis performed after transformation and quality control. The first question, then, is not simply "Can I still use a t-test?" It is: "Am I trying to estimate a mean shift, or detect a broader distributional shift?" [1]
2. What Student's T-Test, Welch's T-Test, and Mann-Whitney U Actually Are
Student's t-test, Welch's t-test, and Mann-Whitney U are related only in the broad sense that all three compare two independent groups. Student's 1908 t-test evaluates whether group means differ under a common-variance model. Welch's 1947 extension asks the same mean-based question, but removes the equal-variance assumption by using group-specific variance terms and adjusted degrees of freedom. Mann and Whitney's 1947 test moves to ranks and asks whether observations from one group tend to be larger than those from the other.
These methods are therefore not interchangeable software options. They target different quantities. Using the wrong test is like measuring liquid volume with a ruler: you still obtain a number, but not the quantity you intended to measure. If your scientific claim is about average abundance or concentration, a rank test is not automatically the cleanest answer. Conversely, if the equal-variance assumption is violated, the pooled-variance t-test can mislead even when the data look broadly well behaved.
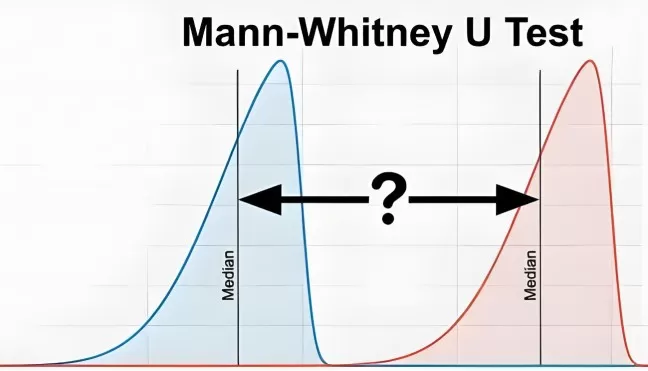
Figure 1. Normal distribution and skewed distribution. Understanding the shape of your data distribution is essential for selecting the appropriate statistical test.
2.1 Mann-Whitney U Test
Mann-Whitney U is a nonparametric test that compares two independent groups by ranking all observations together and testing whether values from one group tend to rank higher than those from the other. It does not assume normality and is robust to outliers, but it tests a different hypothesis than mean-based tests.
3. Why "Non-Normal → Nonparametric" Is a Bad Shortcut
This is why the familiar rule of thumb—"normal → t-test, non-normal → Mann-Whitney"—is too crude for omics. In metabolomics, log transformation is often biologically sensible because many signals are multiplicative and heteroscedastic. However, van den Berg and colleagues showed that log transformation removes heteroscedasticity perfectly only when relative standard deviation is constant, which is rarely true in real datasets; low-abundance features may still remain problematic. In proteomics and multiplexed experiments, the issue is often not just distributional shape but variance structure: technical variation, unbalanced designs, and missing values are common.
This is exactly why the unequal-variance t-test has long been argued to be underused, especially when sample sizes differ. Equally important, Mann-Whitney U is not simply a test of medians. As Hart pointed out, it can also respond to differences in spread and shape. Replacing every "non-normal" feature with a Mann-Whitney U test may therefore swap one assumption problem for a different interpretation problem [2].
4. T-Test vs Welch's T-Test vs Mann-Whitney U: What Biological Question Are You Asking?
Each test answers a different biological question. Student's t-test asks whether the expected mean abundance differs under an equal-variance model. Welch's t-test asks the same mean question, but remains more credible when one group is clearly more variable than the other. Mann-Whitney U asks whether one group tends to rank higher than the other; when distribution shapes are similar, this may behave like a location comparison, but once spread or shape diverges, the interpretation becomes broader. That is why the same molecule may be significant by Welch's t-test but not by Mann-Whitney U, or vice versa, without any calculation being wrong. In many cases, the difference arises because the tests are addressing different inferential questions.
This comparison follows the original formulations of Student, Welch, and Mann-Whitney, together with later clarifications on unequal-variance use and on why rank tests are not automatically median tests.
| Test | Primary Question | When It Is Usually the Better Fit | Natural Biological Wording |
|---|---|---|---|
| Student's t-test | Are the group means different? | Balanced design, similar spread, sensible transformation, no strong evidence of variance inequality | "The mean abundance/concentration differs between groups." |
| Welch's t-test | Are the group means different when variances may differ? | Real-world two-group omics with heteroscedasticity or unequal sample sizes | "The mean differs, allowing for unequal variances." |
| Mann-Whitney U | Do values in one group tend to rank higher, or do distributions differ in location/order? | Strong skew, influential outliers, or a genuine distributional question | "The distributions or rank positions differ between groups." |
5. Why Test Choice Matters So Much in Omics Data Analysis
This distinction matters disproportionately in omics because the data-generating process itself often creates variance asymmetry. In proteomics, DEqMS showed that variance depends on the number of peptides or PSMs supporting protein quantification, meaning that uncertainty is feature-specific rather than constant across the expression matrix. In isobaric-label proteomics, MSstatsTMT highlights multiple conditions, multiple biological and technical replicate runs, unbalanced designs, and missing values as core downstream analysis challenges. In metabolomics, preprocessing choices can change the ranking of biologically important metabolites, and missingness may depend on run day; imputation can increase power, but it can also under- or overestimate effect sizes, while single imputation may underestimate variability and inflate Type I error.
Under those conditions, the same molecule can enter or leave the significant list simply because the statistical test and preprocessing pipeline target different error structures. That is not a software nuisance. It is a biological interpretation problem [3].
6. Examples from Metabolomics, Proteomics, and Spatial Omics
Real studies make this distinction concrete. In the pulmonary nodule study above, targeted assays confirmed higher pyruvate and lactate in stage I lung adenocarcinoma than in benign nodules or healthy controls, while benign and healthy groups were not significantly different. In a setting like this, Welch's t-test is often a better confirmatory choice than the pooled-variance t-test when the disease group is more dispersed, because the biological claim is usually about average concentration after transformation and quality control.
Proteomics offers a different cautionary example. Longitudinal plasma proteome profiling has shown that circulating proteomes are highly individual-specific, and large inflammatory-protein studies document complex variation across many proteins and participants. For an inflammation-related protein with a long upper tail driven by a subset of patients, Mann-Whitney U can be a robust screening tool, but the claim should be framed as a rank or distributional shift, not automatically as mean upregulation. Spatial omics raises a further issue. In oral squamous cell carcinoma, tumor core and leading-edge regions showed distinct transcriptional programs, neighboring cellular compositions, and ligand-receptor interactions. If such data are collapsed into a single average too early, the localized enrichment pattern that matters most biologically may disappear [4].
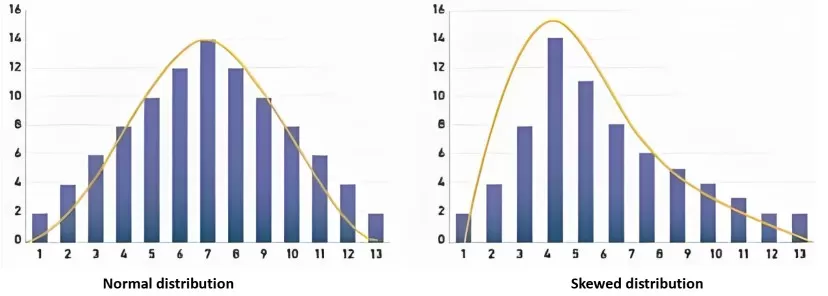
Figure 2. Statistical test selection workflow for omics data analysis. The decision tree helps researchers choose between Student's t-test, Welch's t-test, and Mann-Whitney U based on data characteristics and biological questions.
7. Best Practices for Reporting Two-Group Tests in Omics
The practical takeaway is straightforward. For a simple two-group mean comparison in real proteomics or metabolomics data, Welch's t-test is often a more defensible default than the classical pooled-variance t-test. Mann-Whitney U remains valuable, but it is not a universal replacement. It is most appropriate when the scientific question genuinely concerns ordering or distributional shift, or when extreme values make a rank-based summary more faithful to the biology [5].
In reports, do not stop at the p-value. State the analysis scale, describe the missing-value strategy, show boxplots or violin plots with raw data points, and report an interpretable effect size such as mean difference or fold change. For communication, the most useful visuals are often a three-test comparison table, a boxplot or violin plot that makes unequal variance and outliers visible, and a simple decision tree that begins with the question: "mean difference or distributional difference?" In full omics pipelines, test choice should also be reviewed together with QC, normalization, batch correction, and model structure. Modern frameworks such as limma exist precisely because real omics experiments are small, noisy, and structurally complex.
8. Conclusion: How to Choose the Right Two-Group Test for Your Omics Study
When omics data are not normal, the safest response is not to reach automatically for a nonparametric test. The better response is to clarify the scientific target, inspect the variance structure, and choose the method that matches both the biology and the preprocessing reality. Student's t-test, Welch's t-test, and Mann-Whitney U are all useful precisely because they answer different questions.
Once that distinction is clear, conflicting p-values become easier to interpret: they often mean that the question changed, not just the algorithm. For most real two-group omics workflows—especially those with heteroscedasticity and unequal sample sizes—Welch's t-test is a strong default for mean-based inference. Mann-Whitney U remains essential when the biological story is genuinely about rank or distributional shift. The goal is not to be parametric or nonparametric. The goal is to make a statistical statement that still means the same thing when it reaches the biology section of the paper.
Need Support with Omics Data Analysis?
Choosing the right statistical test is only one part of a reliable omics workflow. MetwareBio provides proteomics, metabolomics, lipidomics, and multi-omics services with comprehensive downstream analysis to help you generate more robust and biologically meaningful results.
Our MetWare Cloud Platform provides integrated bioinformatics analysis tools and expert support for statistical test selection, data preprocessing, and biological interpretation.
Contact us to discuss your project and find the right analysis strategy.
Contact UsReferences
- Yao Y, Wang X, Guan J, Xie C, Zhang H, Yang J, Luo Y, Chen L, Zhao M, Huo B, Yu T, Lu W, Liu Q, Du H, Liu Y, Huang P, Luan T, Liu W, Hu Y. Metabolomic differentiation of benign vs malignant pulmonary nodules with high specificity via high-resolution mass spectrometry analysis of patient sera. Nat Commun. 2023 Apr 24;14(1):2339. doi: 10.1038/s41467-023-37875-1.
- van den Berg RA, Hoefsloot HC, Westerhuis JA, Smilde AK, van der Werf MJ. Centering, scaling, and transformations: improving the biological information content of metabolomics data. BMC Genomics. 2006 Jun 8;7:142. doi: 10.1186/1471-2164-7-142.
- Zhu Y, Orre LM, Zhou Tran Y, Mermelekas G, Johansson HJ, Malyutina A, Anders S, Lehtiö J. DEqMS: A Method for Accurate Variance Estimation in Differential Protein Expression Analysis. Mol Cell Proteomics. 2020 Jun;19(6):1047-1057. doi: 10.1074/mcp.TIR119.001646.
- Hart A. Mann-Whitney test is not just a test of medians: differences in spread can be important. BMJ. 2001 Aug 18;323(7309):391-3. doi: 10.1136/bmj.323.7309.391.
- Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015 Apr 20;43(7):e47. doi: 10.1093/nar/gkv007.


