Not sure whether to use a t-test, Wilcoxon test, ANOVA, or Kruskal-Wallis test for proteomics data? The right choice depends on your study design, sample grouping, and data distribution. This guide helps you understand when each statistical method should be used in differential protein expression analysis, so you can make more reliable and biologically meaningful decisions.
Differential protein expression analysis is a core task in functional proteomics and molecular biology, used to identify proteins that change significantly across biological states, disease conditions, or experimental treatments. The reliability of these findings depends not only on data quality and preprocessing, but also on selecting an appropriate statistical test. In this article, we examine four commonly used methods in proteomics—Student's t-test, the Wilcoxon rank-sum test, analysis of variance (ANOVA), and the Kruskal-Wallis test—covering their principles, assumptions, strengths, limitations, and typical application scenarios. Because these tests are often performed protein by protein in discovery proteomics, P values should generally be interpreted together with fold change and multiple-testing correction, such as the Benjamini-Hochberg false discovery rate (FDR).
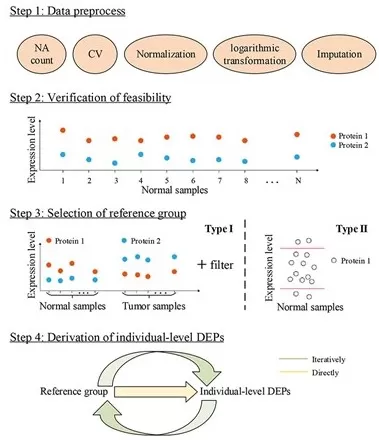
Figure 1. The typical workflow of differential protein expression analysis.
1. Two-Group Statistical Tests in Proteomics
1.1 Student's t-Test in Proteomics
The Student's t-test is a parametric method used to determine whether the means of two groups are significantly different. In proteomics, it is commonly applied when comparing a treatment group with a control group, or when analyzing matched samples such as tumor tissue and paired adjacent normal tissue. Depending on study design, the test can be used either as an independent-samples t-test or as a paired t-test.
t-Test Formula and Principle
For independent samples, it is important to distinguish the classic Student's t-test from Welch's t-test. The classic Student's t-test assumes equal variances and uses a pooled estimate of variance.
t = (X̄₁ − X̄₂) / [sₚ × √(1/n₁ + 1/n₂)]
sₚ² = [((n₁ − 1)s₁²) + ((n₂ − 1)s₂²)] / (n₁ + n₂ − 2)
df = n₁ + n₂ − 2
Welch's t-test does not assume equal variances and is generally preferred when heteroscedasticity is plausible.
t = (X̄₁ − X̄₂) / √(s₁²/n₁ + s₂²/n₂)
df ≈ (s₁²/n₁ + s₂²/n₂)² / [((s₁²/n₁)² / (n₁ − 1)) + ((s₂²/n₂)² / (n₂ − 1))]
Here, X̄₁ and X̄₂ are the sample means, s₁² and s₂² are the sample variances, n₁ and n₂ are the sample sizes, and sₚ² is the pooled variance.
For paired data, the test is based on within-pair differences rather than raw intensities, which makes it especially useful in matched proteomics designs because subject-level variability is partially cancelled out.
t = D̄ / (sD / √n)
df = n − 1
Here, D̄ is the mean of the paired differences, sD is the standard deviation of the paired differences, and n is the number of pairs.
t-Test Assumptions in Proteomics
The t-test assumes independence between observations. For the classic independent-samples Student's t-test, the data within each group should be approximately normal and the group variances should be similar. For Welch's t-test, normality is still important but the equal-variance assumption is relaxed. For a paired t-test, the paired differences should be approximately normal.
t-Test Example in Proteomics
Consider a study comparing the mass-spectrometry-derived abundance of protein X between plasma samples from ten lung cancer patients and ten healthy volunteers after log2 transformation and normalization. A typical workflow would begin with boxplots and formal diagnostic checks such as the Shapiro-Wilk test for normality and Levene's test for variance homogeneity. If the data are approximately normal and variances are similar, the classic independent-samples Student's t-test is an appropriate choice. If variances differ, Welch's t-test provides a safer alternative. For a single prespecified protein, a nominal P value threshold such as 0.05 may be acceptable. In discovery-scale proteomics, however, the result should usually be interpreted with an adjusted P value or FDR together with the log2 fold change so that both statistical confidence and biological effect size are considered.
In a paired design, such as comparing tumor tissue and adjacent non-tumor tissue from the same ten patients, the paired t-test is the correct statistical framework. Because each patient serves as his or her own control, this design can reduce between-subject variation and increase power to detect a true differential protein signal.
t-Test Strengths and Limitations
The t-test has a mature theoretical foundation, is easy to interpret, and is one of the most powerful methods for detecting differences in means when its assumptions are satisfied. It also supports intuitive effect-size reporting through mean differences and confidence intervals. However, it can be sensitive to non-normality in very small sample sets, it is strongly influenced by extreme outliers, and the standard independent-samples version becomes unreliable when heteroscedasticity is ignored.
1.2 Wilcoxon Rank-Sum Test in Proteomics
The Wilcoxon rank-sum test, also known as the Mann-Whitney U test, is a nonparametric alternative for comparing two independent groups. Instead of comparing means, it evaluates whether the two groups occupy different positions in the overall distribution. In many proteomics workflows, it is used when sample sizes are small, data are skewed, or outliers make parametric assumptions difficult to defend.
Wilcoxon Test Formula and Principle
The method pools all observations from both groups, orders them from smallest to largest, assigns ranks, and then calculates the rank sums for the two groups. From these rank sums, the Mann-Whitney U statistic can be computed directly. Under the null hypothesis, the two groups come from the same distribution; for larger samples, U can be converted to a normal approximation.
U₁ = R₁ − n₁(n₁ + 1)/2
U₂ = R₂ − n₂(n₂ + 1)/2
U = min(U₁, U₂)
Here, R₁ and R₂ are the sums of ranks for group 1 and group 2, respectively; n₁ and n₂ are the sample sizes of group 1 and group 2, respectively; U₁ and U₂ are the Mann–Whitney test statistics calculated for the two groups; U is the smaller of U₁ and U₂ and is typically used as the reported test statistic.
Wilcoxon Test Assumptions in Proteomics
The Wilcoxon rank-sum test assumes independence between observations and requires the data to be at least ordinal, meaning they can be ranked. It does not require a specific distributional form, which is a major practical advantage. However, the cleanest interpretation as a location or median shift is obtained when the two group distributions have a similar shape and differ mainly in central tendency.
Wilcoxon Test Example in Proteomics
Imagine comparing normalized abundance values for a target protein between two cell populations, with five biological replicates per group. If one group clearly violates normality, or if boxplots reveal strong skewness or outliers, the Wilcoxon rank-sum test is often more defensible than a standard t-test. A significant result indicates that the distributions differ between the two groups. In reporting, it is often more informative to describe the median and distributional shift rather than forcing a mean-based interpretation.
For matched samples, the corresponding nonparametric method is the Wilcoxon signed-rank test, which evaluates whether the median of the paired differences is zero. This paired version is a useful fallback when a paired t-test is questionable because the distribution of differences is strongly non-normal.
Wilcoxon Test Strengths and Limitations
The Wilcoxon rank-sum test is robust to non-normality, tolerant of outliers, and often safer for very small sample sizes when distributional assumptions are uncertain. At the same time, it generally has lower statistical power than the t-test when the data are in fact approximately normal. It also uses rank information rather than the full quantitative scale, which means some information is lost, and interpretation can become ambiguous when the two groups differ not only in location but also in variance or shape.
(You may also interest at: T-Test vs Welch's T-Test vs Mann–Whitney U)
2. Multi-Group Statistical Tests in Proteomics
2.1 ANOVA for Differential Protein Expression
When a proteomics study involves three or more independent groups, such as control, low-dose treatment, and high-dose treatment, a series of pairwise t-tests is not an ideal starting point. Analysis of variance, or ANOVA, provides a formal parametric framework for testing whether all group means are equal in a single model.
ANOVA Formula and Principle
In one-way ANOVA, the total variability in the data is partitioned into between-group variability and within-group variability. The total sum of squares (SST) reflects overall variation, the between-group sum of squares (SSB) captures variation attributable to group differences, and the within-group sum of squares (SSW) represents random error or individual variation.
SST = SSB + SSW
F = (SSB / (k - 1)) / (SSW / (N - k))
Here, k is the number of groups and N is the total sample size. A large F value indicates that the between-group variation is too large to be explained by random error alone.
ANOVA Assumptions in Proteomics
ANOVA assumes independent observations, approximate normality of residuals within each group, and homogeneity of variances across groups. In practice, variance homogeneity is especially important when group sizes are unbalanced, and Levene's test is often preferred because it is more robust than Bartlett's test when normality is uncertain.
ANOVA Example in Proteomics
Suppose a researcher investigates how a drug affects inflammatory protein Y in cultured cells, with three groups - control, low dose, and high dose - and six biological replicates per group. After log2 transformation and normalization, the analyst first checks normality and variance homogeneity. If the assumptions are reasonable, one-way ANOVA can be performed to test whether at least one group mean differs from the others. A significant ANOVA result, however, does not identify which groups differ. That is why post-hoc multiple comparisons are essential.
Common post-hoc procedures include Tukey's honestly significant difference test for all pairwise comparisons, Dunnett's test when several treatment groups are compared against one control, and Bonferroni-adjusted pairwise tests when a stricter correction is desired. In this way, ANOVA serves as the overall screening step, while the post-hoc analysis pinpoints the specific group contrasts responsible for the signal.
ANOVA can also be extended beyond the one-factor setting. In more complex experimental designs, such as those involving both genotype and treatment, multi-factor ANOVA allows researchers to evaluate not only main effects but also interaction effects, which can be particularly important in mechanistic proteomics studies.
ANOVA Strengths and Limitations
ANOVA offers an efficient and statistically coherent way to analyze multi-group mean differences, avoids the chaos of performing many unstructured pairwise tests, and scales naturally to richer experimental designs. Its main limitations are the same as those of other parametric methods: it is sensitive to strong outliers, depends on distributional assumptions, and requires careful follow-up testing because the omnibus F test alone does not provide biologically actionable pairwise conclusions.
2.2 Kruskal-Wallis Test in Proteomics
The Kruskal-Wallis H test is the nonparametric counterpart of one-way ANOVA. It is used when three or more independent groups need to be compared but the assumptions required for ANOVA are not well supported. In proteomics, this situation often arises when sample sizes are small, abundance distributions are strongly skewed, or outliers remain influential even after standard preprocessing.
Kruskal-Wallis Formula and Principle
The method combines the observations from all groups, ranks them jointly, and calculates the rank sums for each group. The H statistic summarizes how far the group rank profiles deviate from what would be expected if all groups came from the same distribution.
H = [12 / (N(N + 1))] × Σ(Rᵢ² / nᵢ) − 3(N + 1)
Here, N is the total sample size across all groups, nᵢ is the sample size of group i, and Rᵢ is the sum of ranks for group i. When ties are present, a tie-correction factor should be applied. For sufficiently large samples, H is approximately chi-square distributed with k - 1 degrees of freedom.
Kruskal-Wallis Assumptions in Proteomics
Like the Wilcoxon rank-sum test, the Kruskal-Wallis test assumes independent observations and data that can be ranked. Although it does not require normality, interpretation is most straightforward when the group distributions have similar shapes and differ mainly in central tendency rather than in spread or skewness.
Kruskal-Wallis Example in Proteomics
Consider a study measuring the abundance of signaling protein Z across four glioma subtypes, with five samples per subtype. Exploratory analysis may show clear skewness or failed normality diagnostics, making ANOVA difficult to justify. In that setting, the Kruskal-Wallis test provides a more robust omnibus comparison. If the result is significant, the next step is a nonparametric post-hoc procedure such as Dunn's test or the Conover-Iman test, with appropriate multiple-testing correction applied to the pairwise comparisons.
Kruskal-Wallis Strengths and Limitations
The Kruskal-Wallis test is valuable because it relaxes distributional assumptions and performs well for skewed or small-sample data. However, it usually has lower power than ANOVA when parametric assumptions are reasonably satisfied. It also relies only on rank information and may respond to differences in shape as well as differences in central tendency, which means biological interpretation should be made carefully.
3. How to Choose a Statistical Test in Proteomics
In practice, statistical test selection should follow a structured decision path rather than habit or software defaults. The first step is always to clarify the study design: are you comparing two groups or multiple groups, and are the samples independent or paired? For two independent groups, the main decision is usually between a t-test and a Wilcoxon rank-sum test. For paired data, the comparison is typically between a paired t-test and a Wilcoxon signed-rank test. For three or more independent groups, the corresponding choice is usually between ANOVA and Kruskal-Wallis. If the same subjects are measured across three or more conditions or time points, repeated-measures ANOVA or the Friedman test is more appropriate than one-way ANOVA or Kruskal-Wallis.
The second step is diagnostic evaluation of whether parametric assumptions are reasonable. For small sample sizes, normality can be assessed with the Shapiro-Wilk test and by visual inspection using Q-Q plots or histograms. For independent-group comparisons, variance homogeneity can be assessed using Levene's test or Bartlett's test. Outliers should also be reviewed with boxplots or standardized residual approaches, because a single extreme value can materially affect inference.
If the data are approximately normal and variance assumptions are acceptable, parametric methods are usually preferred because they offer higher statistical power and support interpretable effect-size estimates such as mean differences and confidence intervals. If the data are strongly non-normal, contain influential outliers, or show serious heteroscedasticity, nonparametric methods often provide a more robust alternative. That said, mild deviations from normality are not automatically disqualifying, especially in moderately sized samples where parametric methods are known to be reasonably robust. In high-dimensional proteomics, the selected test should also be paired with appropriate multiple-testing control, most commonly Benjamini-Hochberg FDR correction. (You may also interest at: Differential Feature Screening in Omics)
When the sample size is extremely small, such as three replicates per group, normality testing itself has limited power and can create a false sense of security. In those cases, the safer strategy is often to use a nonparametric approach or to make the choice based on strong domain knowledge about the data-generating process rather than on a formal diagnostic test alone.
4. Proteomics Statistical Test Selection Guide
| Method | Best use case in proteomics |
|---|---|
| Student's t-test / Welch's t-test | Two-group comparisons when data are approximately normal; use the paired version for matched samples and Welch's form when variances differ. |
| Wilcoxon rank-sum test | Two-group comparisons when distributions are non-normal, sample sizes are very small, or outliers make a parametric mean comparison unreliable. |
| ANOVA | Three or more independent groups when the goal is to compare means under a parametric framework and follow with appropriate post-hoc tests. |
| Kruskal-Wallis test | Three or more independent groups when ANOVA assumptions are not well supported and a rank-based comparison is more defensible. |
5. Conclusion: Statistical Testing in Proteomics
There is no single best statistical test for every differential protein expression study. The right method depends on the number of groups, whether samples are paired, whether the data are approximately normal, how strongly outliers influence the distribution, and what type of biological conclusion needs to be supported. In proteomics, good statistical practice does not begin with a favorite test. It begins with the study design, continues through diagnostic checking, and ends with an analysis strategy that is both statistically defensible and biologically meaningful. For discovery-scale datasets, that strategy should usually include fold-change interpretation and multiple-testing control rather than relying on uncorrected P values alone.
From Statistical Testing to Biological Insight
Choosing the right statistical test is an essential step in differential protein expression analysis, but reliable conclusions also depend on robust experimental design, high-quality data generation, and thoughtful downstream interpretation. At MetwareBio, we support proteomics, metabolomics, lipidomics, and multi-omics research with integrated analytical workflows that help researchers move from quantitative data to clearer biological insight. If you are planning a differential expression study, our team can help you build a more reliable and efficient research workflow.
Contact Us

