Functional annotation is a critical step in the interpretation of high-throughput omics datasets, bridging raw sequence data with biological meaning. Across genomics, transcriptomics, proteomics, and metabolomics, assigning putative functions to genes or proteins enables identification of biological pathways, regulatory networks, and phenotype associations that underpin complex biological systems. Core resources for functional annotation include the COG/KOG database for orthologous functional grouping, Gene Ontology analysis for hierarchical functional classification, and KEGG pathway enrichment for mapping genes to biochemical and signaling pathways. Each knowledgebase supports distinct aspects of biological interpretation, and the appropriate choice among them, or an integrative application, can substantially influence downstream insights in multi-omics analysis. This guide systematically examines these three annotation resources, providing researchers with evidence-based criteria for optimal utilization in omics data interpretation.
1. FUNCTIONAL ANNOTATION: PRINCIPLES AND APPLICATIONS IN OMICS STUDIES
Functional annotation refers to the process of assigning biological information to genomic elements—genes, proteins, or metabolites—based on sequence similarity, evolutionary conservation, or curated biological knowledge. In the context of multi-omics analysis, functional annotation transforms raw high-throughput data generated from next-generation sequencing and mass spectrometry into interpretable biological features. For example, predicted open reading frames (ORFs) or proteins from a newly sequenced genome can be classified into functional categories, enabling researchers to infer metabolic capabilities or regulatory potential, while transcriptomic or proteomic datasets often require functional annotation to link differential expression to biological processes or cellular components. Functional annotation thus acts as a bridge from large, high-dimensional molecular datasets to biologically meaningful interpretations that support hypothesis generation and experimental validation.
The importance of functional annotation is underscored by the complexity of biological systems. High-throughput technologies routinely generate data with tens of thousands of features, and extracting biologically actionable insights requires mapping these features to known functions or pathways. Without functional annotation, researchers are left with lists of genes or proteins that lack context, hindering efforts to understand molecular mechanisms of disease, identify drug targets, or explore phenotypic variation. Tools and frameworks such as the COG KOG database, Gene Ontology analysis, and KEGG pathway enrichment provide structured vocabularies and pathway maps that support interpretation of omics data. The integration of annotations across these resources enhances the resolution of biological interpretation, enabling systems-level insights into molecular interactions and phenotypes. Moreover, functional annotation is increasingly vital in multi-omics strategies that combine multiple data layers to reveal networks of regulation and function across genomic, transcriptomic, proteomic, and metabolomic levels.
2. COG/KOG DATABASE: ORTHOLOG-BASED FUNCTIONAL CLASSIFICATION
Functional annotation with the COG/KOG database relies on principles of evolutionary biology and homology inference. The Clusters of Orthologous Groups (COG) database, originally developed for prokaryotic genomes, and its eukaryotic counterpart, KOG (Eukaryotic Orthologous Groups), offer a structured framework for classifying proteins based on evolutionary relationships. These databases are constructed through systematic comparison of protein sequences from fully sequenced genomes, grouping highly homologous proteins into clusters that represent orthologous relationships across species (Tatusov et al., 2003). Orthologous groups are divided into 26 functional categories covering key cellular processes such as information storage and processing, metabolism, and general cellular functions, providing a high-level functional overview that is especially useful for newly sequenced genomes or non-model organisms with limited annotation. The COG collection originally encompassed approximately 138,458 proteins forming 4,873 clusters, covering 75% of predicted proteins from 66 prokaryotic genomes, while KOG included 4,852 clusters derived from seven eukaryotic genomes, including human, mouse, and other model organisms (Tatusov et al., 2003). Recent updates have further expanded COG coverage to thousands of bacterial and archaeal genomes, enhancing the depth and reliability of functional annotation for prokaryotic species (Galperin et al., 2025).

Figure 1. The 26 COG functional categories in NCBI COG Database. Image adapted from NCBI COG Database.
2.1 COG/KOG Annotation and Enrichment Workflow
A typical workflow using the COG/KOG database begins with predicted protein sequences derived from genomic or proteomic data. These sequences are aligned against a reference orthologous group database such as COG/KOG using sequence similarity search tools (e.g., DIAMOND, BLAST). Matches are assigned to specific COG/KOG identifiers, which link the query sequences to functional categories. Tools like eggNOG-mapper (based on extended orthologous group databases) automate this process, enabling scalable annotation of large omics datasets. Enrichment analysis then assesses whether specific COG/KOG functional categories are over-represented in a subset of proteins or differential gene sets relative to a background distribution, typically using hypergeometric tests or related statistical approaches.
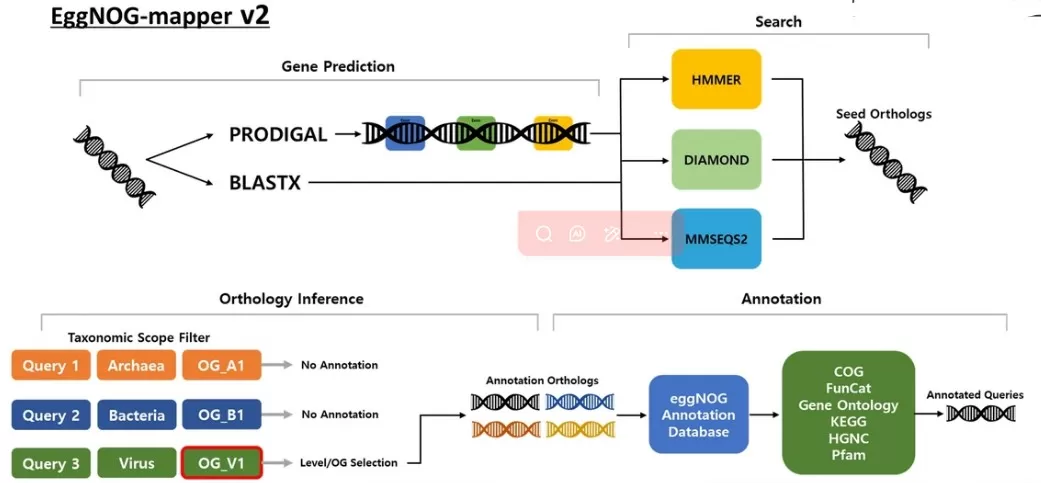
Figure 2. The workflow of EggNOG-mapper v2 consists of gene prediction, search, orthology inference, and annotation stages. Image adapted from Kim et al. (2024), Journal of Translational Medicine, licensed under CC BY 4.0.
2.2 Advantages, Limitations, and Applications of COG/KOG
The strengths of COG/KOG-based annotation lie in its orthology focus and evolutionary context. It provides a broad overview of functional class distributions and supports comparative analyses across prokaryotic and eukaryotic genomes. Because the classifications are coarse-grained, they can highlight overarching functional themes in large datasets. However, limitations include relatively lower resolution for detailed functional nuance compared to ontology-based systems like Gene Ontology, and limited coverage for organisms without well-characterized orthologs. COG/KOG annotation is most effective for microbial functional profiling, comparative genomics, and evolutionary studies, and forms a valuable component of multi-omics annotation pipelines that combine multiple knowledgebases for deeper biological insight.
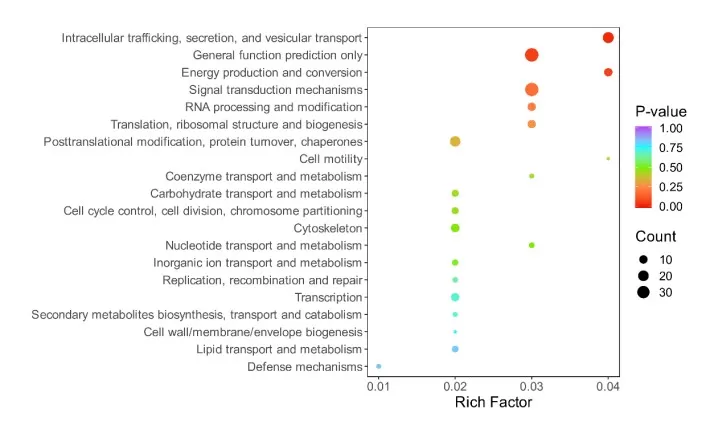
Figure 3. KOG Enrichment Analysis Bubble Plot.
3. GENE ONTOLOGY (GO): HIERARCHICAL FUNCTIONAL ANNOTATION
Functional interpretation in omics often relies on structured vocabularies, and Gene Ontology (GO) provides a comprehensive hierarchical framework for classifying gene products by their functions (Gene Ontology Consortium, 2021). GO annotations categorize genes or proteins into three ontologies: Cellular Component (the subcellular structures where gene products act), Molecular Function (activities at the molecular level, such as catalytic or binding activities), and Biological Process (broader pathways or processes involving multiple molecular activities). Each term within the ontology represents a defined biological concept, and the annotations describe how gene products contribute to these concepts based on experimental evidence or computational inference. GO's structured, controlled vocabulary enables uniform functional annotation across species and supports nuanced interpretation of biological roles and processes.
3.1 GO Annotation and Enrichment Workflow
The GO functional annotation workflow using Gene Ontology (GO) begins with mapping gene or protein identifiers to GO terms. Widely used tools for high-throughput GO annotation include Ensembl BioMart, UniProt GO mapping, PANTHER, DAVID, g:Profiler, ShinyGO, and WebGestalt, which assign GO terms based on sequence similarity, conserved domains, or curated functional associations. Following annotation, GO analysis can quantify the distribution of GO terms across a dataset and compare their representation between experimental groups. Enrichment analysis is then performed to identify terms that are statistically over-represented, using commonly applied tools such as topGO, clusterProfiler, GOstats, and BiNGO. Statistical approaches include over-representation analysis (ORA) and gene set enrichment analysis (GSEA), which determine whether specific GO terms appear more frequently than expected by chance in a target gene or protein list (Muley, 2025). The results highlight biological processes, molecular functions, or cellular components most relevant to the experimental condition, providing structured insights for interpretation of omics datasets.
 Functional Annotation Bar Chart_1776322744_WNo_993d467.webp)
Figure 4. Gene Ontology (GO) Functional Annotation Bar Chart.
3.2 Advantages, Limitations, and Applications of GO Analysis
The primary advantage of Gene Ontology (GO) lies in its detailed granularity and standardized hierarchical structure, which enables precise functional interpretation across diverse biological systems. GO annotations allow researchers to identify specific biological processes, molecular functions, and cellular components, making them particularly valuable for interpreting differential expression in transcriptomic and proteomic datasets. However, several challenges accompany GO-based analyses. Redundancy and hierarchical complexity can complicate interpretation, especially when multiple related terms appear significant, and annotation coverage varies across organisms and data types, with non-model species often exhibiting limited GO representation. Moreover, benchmarking studies have highlighted substantial variability among GO enrichment tools: different software packages can produce divergent results even when analyzing identical input data with the same underlying GO resource (de Oliveira et al., 2026). This variability emphasizes the importance of carefully selecting appropriate tools and validation strategies. Despite these limitations, GO remains the most widely adopted ontology for functional annotation and enrichment in omics workflows, particularly when used in combination with other annotation systems in multi-omics analyses.
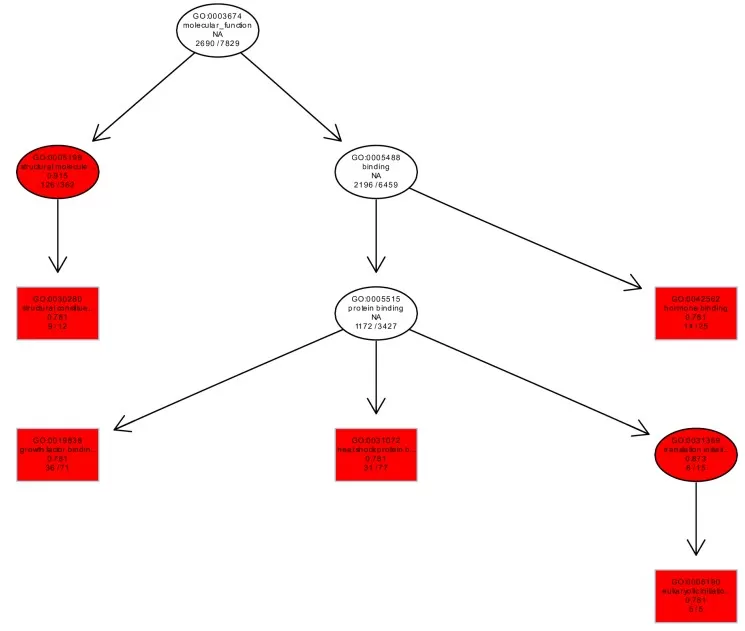
Figure 5. GO Enrichment Directed Acyclic Graph.
4. KEGG FUNCTIONAL ANNOTATION: FROM SINGLE FEATURES TO SYSTEM-LEVEL PATHWAYS
KEGG (Kyoto Encyclopedia of Genes and Genomes) is a comprehensive knowledgebase that integrates genomic information with systemic functional contexts such as biochemical pathways, molecular interactions, and environmental information processing. At its core is the KEGG Orthology (KO) system, a structured classification of genes and proteins into functional ortholog groups linked to pathway maps, functional hierarchies, and biochemical modules. KO identifiers (K numbers) connect genes to defined biological systems, enabling mapping of omics features onto pathways such as metabolism, signal transduction, and disease pathways. The KEGG database thus provides a pathway-centric view of functional annotation that supports interpretation of biological systems at a systems level.
4.1 KEGG Annotation and Enrichment Workflow
KEGG annotation begins with assigning KO membership to gene or protein sequences, typically via tools such as BlastKOALA, GhostKOALA, or KofamKOALA that search query sequences against KEGG reference data. Once assigned, KO terms provide direct linkage to curated pathways and modules within the KEGG database. KEGG pathway enrichment analysis statistically evaluates whether a set of genes or proteins is over-represented in specific pathways compared with a background distribution, highlighting biological systems significantly impacted under experimental conditions. This enrichment often employs hypergeometric testing similar to GO enrichment but yields interpretation focused on pathways rather than individual functional terms. KEGG mapping also enables visual representation of pathways with highlighted gene/protein hits, facilitating intuitive interpretation of complex results.
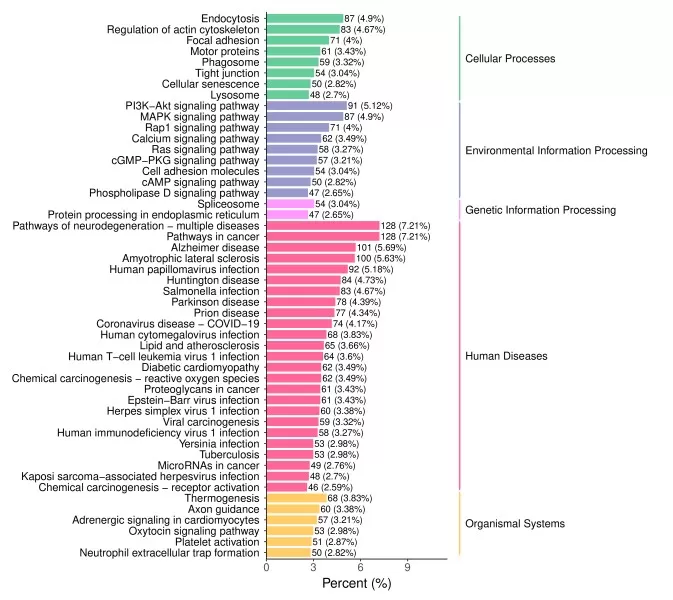
Figure 6. Functional Annotation Bar Chart.
4.2 Advantages, Limitations, and Applications of KEGG Pathway Analysis
KEGG's strength lies in its curated integration of functional orthologs with pathway knowledge, providing a comprehensive context for interpreting biochemical and cellular systems. Pathway maps offer insight into the interplay between metabolic routes, signaling cascades, and functional modules, which is particularly valuable in metabolomics and integrative multi-omics studies. However, limitations include gaps in pathway coverage for some non-model organisms and the static nature of pathway representations that may not capture dynamic regulatory states. Despite these limitations, KEGG pathway annotation and enrichment analysis are widely used in transcriptomics, proteomics, and metabolomics workflows where understanding systemic biological changes is essential.
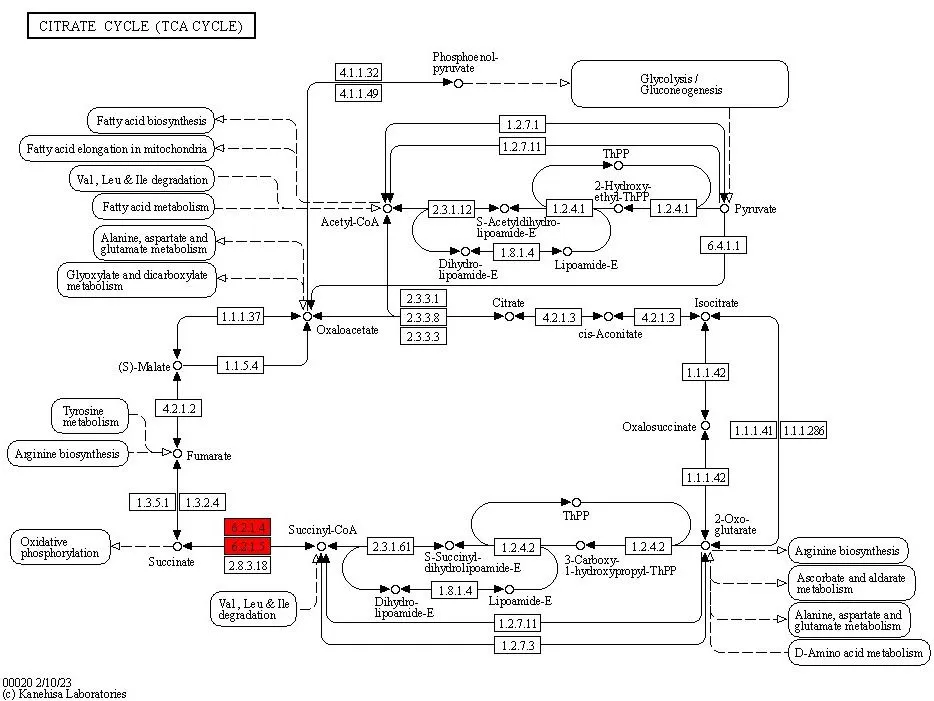
Figure 7. KEGG Pathway Map of Differentially Expressed Proteins.
5. COG/KOG VS GO VS KEGG: SELECTING THE RIGHT FUNCTIONAL ANNOTATION STRATEGY
Understanding the relative strengths and limitations of the COG KOG database, Gene Ontology analysis, and KEGG pathway enrichment is essential for selecting appropriate functional annotation strategies. These resources vary in scope, granularity, and biological context: the COG/KOG approach emphasizes evolutionary conservation and broad functional categories; GO provides detailed hierarchical annotations of biological roles; and KEGG focuses on pathway structures that connect molecular functions with systemic biological processes. Integrative use of these resources can leverage the orthology-based context of COG/KOG with GO's functional detail and KEGG's pathway framework to support more comprehensive interpretation of omics data. A summary comparison is shown below:
| Feature | COG/KOG | GO | KEGG |
|---|---|---|---|
| Core Principle | Orthologous clustering based on sequence homology | Structured ontology with hierarchical terms | Manually curated pathway maps and molecular networks |
| Output Type | Functional category assignment (25 classes) | Term assignments across 3 ontologies | Pathway maps and KO identifiers |
| Species Coverage | Prokaryotes (COG); limited eukaryotes (KOG) | All species (ontology-based) | Primarily model organisms with orthology mapping |
| Update Frequency | Infrequent; stable resource | Regular updates with versioning | Regular updates (monthly) |
| Best Application | Comparative genomics; novel genome annotation | Functional enrichment; species-agnostic analysis | Pathway mapping; metabolic and signaling networks |
| Key Limitation | Incomplete eukaryotic coverage | Tool-dependent result variability | Non-model species coverage gaps |
Each annotation strategy addresses different interpretive needs within multi-omics analysis. COG/KOG is particularly useful for broad comparisons across genomes or functional profiling of large gene sets, while GO is optimal for detailed characterization of gene or protein function in specific biological contexts. KEGG excels in depicting interactions among molecular entities within pathways, making it particularly valuable for interpreting metabolic and signaling changes. In practice, many workflows combine these resources, using COG/KOG to categorize sequences, GO to detail functional roles, and KEGG to contextualize those roles within pathways. This integrative strategy enhances the biological interpretability of high-throughput omics datasets, supporting comprehensive multi-omics analysis that spans from individual gene function to systems-level mechanisms.
6. BEST PRACTICES FOR FUNCTIONAL ANNOTATION IN OMICS STUDIES
Effective functional annotation in omics research requires thoughtful preparation, well-designed annotation strategies, and robust integration of results to ensure biological insights are meaningful and reproducible. Integrating information from diverse annotation resources within a multi-omics analysis framework maximizes interpretive depth and supports rigorous scientific conclusions.
6.1 Data Preparation and Quality Control
Accurate functional annotation depends on rigorous data preprocessing and quality control. Key steps include missing value imputation to address incomplete entries in high-throughput omics datasets, data normalization to correct for technical variation and ensure comparability across samples, and batch effect correction to minimize systematic differences between experimental runs. Input datasets should also be curated to maintain consistent gene or protein identifiers and remove duplicate entries. Quality control assessments, including Principal Component Analysis (PCA) or hierarchical clustering, help detect outliers and evaluate overall data structure. Implementing these preprocessing and QC procedures establishes a reliable foundation for mapping features to functional categories and performing downstream GO, COG/KOG, or KEGG enrichment analysis, enhancing interpretability and reproducibility in multi-omics studies.
6.2 Annotation Strategy
Annotation strategies should align with research hypotheses and data types. A single-database approach may suffice for specific tasks (e.g., broad functional profiling with a COG/KOG database), but integrating multiple resources (e.g., GO terms with KEGG pathways) offers a richer biological context. Overrepresentation analysis (ORA) remains the most widely used enrichment method; however, benchmarking studies reveal substantial heterogeneity among ORA tools in terms of biological informativeness and result specificity (de Oliveira et al., 2026). Statistical considerations include appropriate background selection (genome-wide or transcriptome-wide), multiple testing correction (Benjamini-Hochberg FDR preferred), and significance threshold selection (commonly FDR < 0.05). Multi-omics studies often require harmonization of annotation across different data layers, which may involve cross-mapping gene IDs or reconciling database versions. Enrichment results should be interpreted with attention to term specificity—broad, generic terms often provide less biological insight than more specific, granular annotations.
6.3 Integrative Analysis Tips
Integrative multi-omics analysis provides a comprehensive view of biological systems but presents challenges in data alignment, scaling, and interpretation. Combining functional annotation results across different omics layers—such as transcriptomics and metabolomics—can enhance biological insights; however, it requires careful data normalization and attention to technical variation between datasets. Effective visualization strategies, including network diagrams for GO terms, pathway maps for KEGG results, and functional category bar plots for COG/KOG analyses, support intuitive interpretation and clear communication of results. Comprehensive reporting with structured tables and figures further facilitates hypothesis validation and downstream experimental planning. Ensuring reproducibility requires documenting database versions (e.g., GO release date, KEGG version), annotation tool settings, and statistical thresholds applied. Adhering to these best practices strengthens the interpretability, reliability, and biomedical relevance of functional annotation outcomes, making the insights derived from multi-omics datasets robust and actionable for research and drug discovery applications.
7. CONCLUSION AND TRENDS OF MULTI-OMICS FUNCTIONAL ANNOTATION
Functional annotation is indispensable for interpreting omics and multi-omics data, enabling translation of raw molecular features into biological meaning. The COG KOG database provides an orthology-based foundation for broad functional classification, Gene Ontology analysis delivers detailed functional characterization, and KEGG pathway enrichment contextualizes molecular changes within systemic pathways. Each resource contributes distinct yet complementary perspectives that support comprehensive biological interpretation.
Emerging trends in omics research emphasize integration across multiple annotation resources to capture complex biological relationships that single databases alone cannot resolve. Combining orthology-based classification with hierarchical functional annotation and pathway context improves the robustness of biological insights from high-throughput data. As multi-omics approaches become more prevalent, the integration of annotation results across genomic, transcriptomic, proteomic, and metabolomic layers will continue to enhance our understanding of molecular mechanisms and disease processes.
Looking ahead, machine learning and artificial intelligence hold promise for advancing functional annotation and enrichment analysis. AI-driven models are being developed to predict functional annotations from sequence data with high accuracy, addressing current limitations in manual curation and improving coverage for non-model organisms (e.g., transformer-based GO prediction models). Continued development of integrative algorithms that combine statistical enrichment with network inference and predictive modeling will further expand the interpretive power of functional annotation in multi-omics research. These innovations are poised to accelerate discoveries in systems biology, precision medicine, and drug development.
MetwareBio: Turn Functional Annotation Into Actionable Multi-Omics Insights
At MetwareBio, we combine functional annotation with integrated multi-omics analysis to help researchers move from feature lists to biological interpretation. By leveraging tools such as COG/KOG, GO, and KEGG within standardized bioinformatics workflows, we support clearer pathway discovery, functional insight, and data-driven decision-making across proteomics, metabolomics, and other omics studies.
Contact UsReferences
- Tatusov, R. L., Fedorova, N. D., Jackson, J. D., Jacobs, A. R., Kiryutin, B., Koonin, E. V., Krylov, D. M., Mazumder, R., Mekhedov, S. L., Nikolskaya, A. N., Rao, B. S., Smirnov, S., Sverdlov, A. V., Vasudevan, S., Wolf, Y. I., Yin, J. J., & Natale, D. A. (2003). The COG database: an updated version includes eukaryotes. BMC Bioinformatics, 4, 41. https://doi.org/10.1186/1471-2105-4-41
- Galperin, M. Y., Vera Alvarez, R., Karamycheva, S., Makarova, K. S., Wolf, Y. I., Landsman, D., & Koonin, E. V. (2025). COG database update 2024. Nucleic Acids Research, 53(D1), D356–D363. https://doi.org/10.1093/nar/gkae983
- Kim, C., Pongpanich, M., & Porntaveetus, T. (2024). Unraveling metagenomics through long-read sequencing: a comprehensive review. Journal of Translational Medicine, 22(1), 111. https://doi.org/10.1186/s12967-024-04917-1
- Gene Ontology Consortium (2021). The Gene Ontology resource: enriching a GOld mine. Nucleic Acids Research, 49(D1), D325–D334. https://doi.org/10.1093/nar/gkaa1113
- Muley, V. Y. (2025). Functional Insights Through Gene Ontology, Disease Ontology, and KEGG Pathway Enrichment. Methods in Molecular Biology, 2927, 75–98. https://doi.org/10.1007/978-1-0716-4546-8_4
- de Oliveira, F. H. S., Gomes, F. A., & Feltes, B. C. (2026). Benchmarking multiple gene ontology enrichment tools reveals high biological significance, ranking, and stringency heterogeneity among datasets. Frontiers in Bioinformatics, 6, 1755664. https://doi.org/10.3389/fbinf.2026.1755664


