In omics research, "p<0.05" is often treated as the gold standard for statistical significance. But is it really enough? This article explores why professional omics reports must go beyond raw p-values, and how adjusted p-values, FDR, and q-values provide more reliable frameworks for discovery-driven science.
1. WHY P<0.05 IS NOT ENOUGH IN OMICS
Reporting only "p<0.05" in an omics study is a bit like running muddy water through a single coarse filter and then declaring it safe to drink. A raw p-value is useful, but it is only a first-pass signal. It tells you whether one observed difference looks surprising under the null. It does not tell you what happens when that same test is repeated across hundreds, thousands, or even tens of thousands of features.
That is why the p-value vs FDR omics discussion is not just about terminology. It is about the cost of discovery. How many false positives are you willing to carry into downstream validation, internal decision-making, reviewer scrutiny, or client reporting? Multiple-testing correction works like a multi-stage purification system. Adjusted p-values, FDR, and q-values are different ways of describing how much uncertainty remains after that filtering step. The threshold you choose is therefore not just a technical parameter. It reflects your tolerance for risk. In omics, where the aim is often to discover candidate molecules rather than test a single predefined hypothesis, that distinction is essential rather than optional [1].
2. THE REAL PROBLEM IS NOT ONE TEST BUT THOUSANDS
The real challenge in omics is not that p-values are wrong. It is that they are local answers to a global problem. If a study tests 10,000 proteins using a nominal cutoff of p<0.05, chance alone can produce a large number of apparently significant results. In discovery proteomics, this is not a hypothetical concern. Published FFPE lung cancer proteomics studies have already reported more than 8,000 quantified proteins and over 14,000 phosphosites, and large-scale DIA workflows were developed precisely because proteomics datasets have outgrown the scale that older statistical habits were built for [2].
Once the number of tests becomes large, nominal significance becomes increasingly disconnected from shortlist credibility. A seemingly rich discovery list may in fact contain a mixture of true biology and random noise. That is the core of the multiple testing omics problem. A professional report should not stop at saying which features passed a nominal cutoff. It should also indicate how much error is expected in the set of features being carried forward. Without that second layer of interpretation, the word "significant" often means far less than it appears to mean.
3. P-VALUE, ADJUSTED P-VALUE, FDR, AND Q-VALUE: FOUR DIFFERENT RISK LANGUAGES
These four terms are related, but they are not interchangeable. A p-value is the raw statistical evidence for one feature in one test. An adjusted p-value is the multiplicity-corrected version of that feature-level evidence, used to assess whether the feature remains significant after accounting for many parallel tests. FDR, by contrast, is not a property of one feature alone. It describes the expected proportion of false positives within the final set of declared discoveries. The q-value goes one step further: it gives each feature the minimum FDR threshold at which that feature would be called significant [3].
That makes q-values especially useful when you want flexible, tiered prioritization rather than a rigid one-cutoff output. In practical terms, these four measures speak to evidence at different levels: one test, one corrected feature, one shortlisted set, and one feature's minimum acceptable discovery risk. That is why mature omics differential analysis statistics reporting should explain not only the cutoff itself, but also the level of interpretation behind it.
Table 1. Decision-oriented interpretation of p-value, adjusted p-value, FDR, and q-value.
| Measure | Level | Interpretation | Use Case |
|---|---|---|---|
| p-value | One test | Probability of observing this result under the null hypothesis | Initial screening, single hypothesis testing |
| Adjusted p-value | One corrected feature | p-value after multiple testing correction (e.g., Bonferroni, Benjamini-Hochberg) | Conservative feature selection, family-wise error control |
| FDR | Entire discovery set | Expected proportion of false positives among declared discoveries | Discovery-oriented studies, biomarker candidate lists |
| q-value | One feature's minimum FDR | Minimum FDR threshold at which this feature would be significant | Tiered prioritization, flexible significance thresholds |
4. WHY THIS MATTERS ESPECIALLY IN OMICS, AND WHY THRESHOLDS SHOULD CHANGE BY PROJECT STAGE
Omics is not simply classical statistics performed on a larger dataset. It is a different decision environment. In traditional hypothesis testing, the goal may be to evaluate one predefined question. In discovery omics, the goal is usually to generate and rank a candidate list under uncertainty. That changes what significance thresholds are supposed to do.
In early exploratory work, a somewhat broader false-positive margin may be acceptable if the results will later be filtered by pathway consistency, orthogonal evidence, or targeted follow-up. In that context, FDR<0.10 may be reasonable. As the project moves into shortlist refinement, the threshold should become stricter, and FDR<0.05 combined with effect-size and data-quality criteria often becomes more appropriate. Before a feature is taken into validation, the standard should tighten again, because each false positive now consumes real budget, real samples, and real effort.
This is not inconsistency. It is good study design. Threshold choice should reflect where the project sits on the spectrum between exploration and commitment. That is why omics reports should explain threshold strategy in the context of study phase, downstream use, and risk tolerance rather than implying that one universal cutoff is always correct [1].
A simple stage-based framework is shown below. Exact cutoffs may vary by platform and dataset quality, but the underlying logic remains broadly applicable [4].
Table 2. Stage-based threshold logic for omics discovery projects.
| Project Stage | Goal | Suggested FDR | Additional Criteria |
|---|---|---|---|
| Exploratory Discovery | Generate broad candidate list | <0.10 | Effect size, pathway consistency |
| Shortlist Refinement | Prioritize top candidates | <0.05 | Effect size, data quality, reproducibility |
| Validation Preparation | Select features for orthogonal validation | <0.01 | Effect size, biological relevance, sample availability |
5. PROTEOMICS EXAMPLE: WHY ADJUSTED P VALUE IN PROTEOMICS CHANGES THE SHORTLIST
Proteomics is perhaps the clearest context in which adjusted p value in proteomics becomes a reporting necessity rather than a cosmetic refinement. Modern mass spectrometry workflows now handle very large and complex DIA datasets, and published tumor proteomics studies have already reached depths of more than 8,000 quantified proteins. At that scale, a list based only on raw p-values can easily look much larger—and much more convincing—than it really is.
The exact number of disappearing hits will vary by study design, variance, sample size, and missingness, but the overall pattern is predictable: once multiplicity is taken into account, many nominal hits drop out. That should not be seen as a statistical failure. It is a gain in honesty. A shortlist based on adjusted p-values or FDR is more credible because it better reflects the expected error burden across the entire discovery set. This matters especially in proteomics, where missing values and technical variation are common and can destabilize nominal significance if not handled carefully. For experimental teams, reviewers, and clients alike, corrected significance is therefore much closer to a usable decision framework than raw p<0.05 alone [5].
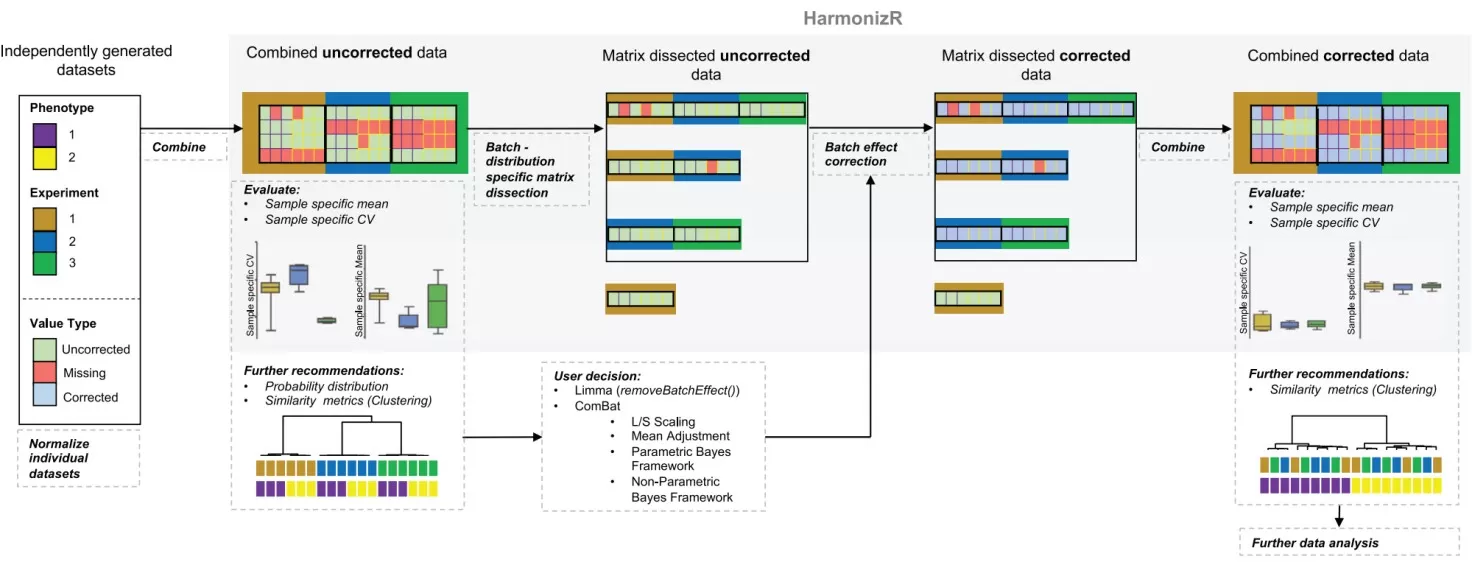
Figure 1. The HarmonizR operation principle for batch effect reduction across independent proteomic studies. Image adapted from Voß et al., 2022, Nature Communications, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
6. METABOLOMICS EXAMPLE: WHY Q VALUE METABOLOMICS IS SO USEFUL FOR TIERED OUTPUT
Metabolomics typically involves fewer confidently identified molecules than deep proteomics, so the gap between raw p-values and corrected values may be less dramatic. But it is never trivial. Statistical workflows for metabolic phenotyping explicitly include normalization, multivariate analysis, candidate biomarker selection, validation, and multiple-testing correction, because even a few hundred features can produce misleading nominal hits if interpreted casually [6].
Representative paired tissue studies, such as the clear cell renal cell carcinoma atlas built from 138 matched tumor and normal tissue pairs, show extensive pathway-level remodeling alongside transcriptome–metabolome discordance. That is exactly the kind of setting in which q value metabolomics becomes particularly useful. Because the q-value gives each feature its own minimum acceptable FDR level, it naturally supports tiered reporting: a strict core set for immediate validation, a medium-confidence set for pathway interpretation, and a broader exploratory set for future follow-up.
That is often more informative than forcing every feature into a binary yes-or-no decision at one arbitrary threshold. In metabolomics, where discovery and interpretation are tightly linked, tiered output is often both more honest and more actionable than a single-cutoff result.
7. CONCLUSION: Choose Thresholds by Discovery Risk, Not Habit
Truly professional omics reporting is not just about calculating p-values and exporting a differential list. It is about translating statistical evidence into an actionable discovery strategy. That translation requires corrected significance, transparent threshold logic, and a prioritization framework that reflects project stage, data quality, and validation cost.
Whether the platform is proteomics, metabolomics, or spatial multi-omics, the strongest report does more than say what is significant. It distinguishes what is credible now, what is promising but still exploratory, and what should wait for stronger evidence. That is increasingly the standard expected by sophisticated clients, reviewers, and collaborators. The real value of omics analysis, therefore, is not in stopping at "p<0.05," but in helping people understand what level of discovery risk they are accepting, why they are accepting it, and which candidates are truly worth validating next [4].
Turn Statistical Significance into Better Omics Decisions
If you need more than a simple p<0.05 cutoff, MetwareBio provides proteomics, metabolomics, lipidomics, and multi-omics services with integrated statistical analysis and clear result interpretation. Explore our omics solutions to build more reliable candidate lists and move forward with greater confidence in downstream validation.
Contact UsRead More on Omics Differential Analysis
- Differential Feature Screening in Omics: Why the Best Candidate Is Not Always the One with the Smallest p-Value
- FDR Control in Proteomics: Principles, Calculation Methods, and Threshold Selection
- Statistical Tests for Differential Protein Expression in Proteomics
- Volcano Plots in Metabolomics & Proteomics: Interpretation, Cutoffs, and Best Practices
References
- Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society: Series B. 1995;57(1):289–300.
- Comprehensive micro-scaled proteome and phosphoproteome characterization of archived retrospective cancer repositories. Nature Communications. 2021;12:3576.
- Storey JD, Tibshirani R. Statistical significance for genomewide studies. Proceedings of the National Academy of Sciences. 2003;100(16):9440–9445.
- Kohler D, Staniak M, Yu F, Nesvizhskii AI, Vitek O. An MSstats workflow for detecting differentially abundant proteins in large-scale data-independent acquisition mass spectrometry experiments with FragPipe processing. Nature Protocols. 2024;19:2915–2938.
- Voß H, et al. HarmonizR enables data harmonization across independent proteomic datasets with appropriate handling of missing values. Nature Communications. 2022;13:3523.
- Statistical analysis in metabolic phenotyping. Nature Protocols. 2022;16:4299–4326.


