For years, metabolomics researchers have had to work around a familiar trade-off. Untargeted metabolomics provides broad discovery power, but often falls short in quantitative accuracy and reproducibility. Targeted metabolomics, by contrast, enables precise measurement of predefined metabolites, yet cannot capture unexpected or novel compounds. This long-standing gap between coverage and accuracy helped drive the rise of widely targeted metabolomics. By combining high-resolution LC-MS/MS in DDA mode for metabolite discovery and annotation with triple quadrupole mass spectrometry in MRM mode for sensitive, reproducible quantification, widely targeted metabolomics offers a practical balance between breadth and rigor. This article explores how the workflow operates, why it matters, and where it is being applied across plant science and biomedical research.
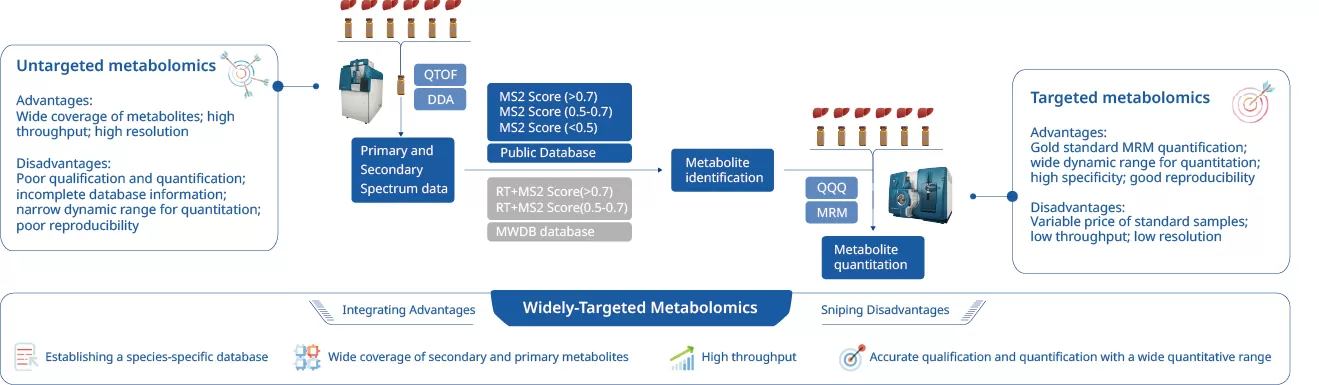
Figure 1. Conceptual Comparison of Untargeted, Targeted, and Widely-Targeted Metabolomics.
1. What Is Widely-Targeted Metabolomics?
Widely-targeted metabolomics (also referred to as large-scale targeted metabolomics or pseudo-targeted metabolomics) is a hybrid analytical strategy that combines the discovery-oriented capabilities of untargeted metabolomics with the quantitative precision of targeted metabolomics within a single integrated platform. Unlike classical untargeted workflows that rely on semi-quantitative peak area comparisons, or targeted panels limited to dozens or hundreds of preselected compounds, widely-targeted metabolomics can simultaneously profile hundreds to thousands of metabolites with genuine quantitative accuracy.
The approach was pioneered by Chen et al. (2013) in Molecular Plant, who developed a novel integrated method for large-scale detection, identification, and quantification of widely targeted metabolites using LC-MS/MS. Since then, the methodology has been refined and expanded across diverse biological matrices — from plant tissues and food commodities to biofluids such as plasma, serum, and urine — establishing itself as a mainstream option for researchers who need both coverage and confidence.
At its core, widely-targeted metabolomics leverages a proprietary database architecture that stores dual sets of spectral information for each metabolite: high-resolution parameters (accurate mass, MS/MS fragmentation patterns, retention time) acquired on instruments like QTOF or Orbitrap, alongside low-resolution but highly sensitive MRM transition parameters (precursor ion Q1, product ion Q3, declustering potential DP, collision energy CE, retention time) optimized for triple quadrupole platforms. This dual-parameter database is the key innovation that enables the seamless transition from qualitative discovery (DDA) to targeted quantification (MRM).
The term "widely-targeted" reflects precisely this design philosophy: it is "targeted" because every detected compound is quantified using the rigorous MRM mode, yet it is "widely" because the target list is not preconstrained — it is dynamically generated from the DDA discovery phase, allowing the method to adapt to each sample type's unique metabolic landscape.
2. How Widely-Targeted Metabolomics Workflow Operates?
The widely-targeted metabolomics workflow consists of five sequential stages that transform raw biological samples into high-quality quantitative metabolic datasets. Each stage builds upon the previous one, creating a pipeline that maximizes both metabolite coverage and data reliability.
Step 1: Sample Preparation and Metabolite Extraction
Upon receipt, samples undergo matrix-adapted metabolite extraction tailored to the specific sample type. The extraction protocol typically involves rapid quenching of enzymatic activity (using liquid nitrogen flash-freezing for plant samples or cold solvent systems for biofluids), followed by homogenization and extraction with appropriate solvent mixtures (e.g., methanol-water-chloroform for polar and lipophilic metabolite recovery). Quality control (QC) samples are prepared at this stage by pooling equal aliquots from all study samples; these pooled QC samples will be used throughout the analytical run to monitor system stability and enable downstream data normalization.
Each processed sample yields a single tube of extracted metabolites ready for LC-MS/MS analysis. Proper sample preparation at this stage is critical — inadequate extraction leads to metabolite degradation or biased recovery, while well-executed preparation ensures reproducible results across hundreds or thousands of samples.
Step 2: DDA-Based Metabolite Discovery on High-Resolution MS
A portion of the pooled QC extract is subjected to chromatographic separation (typically using ultra-high-performance liquid chromatography, UPLC) coupled to a high-resolution mass spectrometer such as a quadrupole time-of-flight (QTOF) or Orbitrap instrument. The MS operates in data-dependent acquisition (DDA) mode, which performs a survey scan (MS1) to detect all ionizable metabolite features, then automatically selects the most intense precursor ions for fragmentation (MS2) to generate characteristic product ion spectra.
During this stage, three critical pieces of information are captured for each detectable metabolite:
- Accurate mass (MS1): The precise mass-to-charge ratio (m/z) enabling elemental composition prediction (typically < 5 ppm error)
- MS/MS fragmentation pattern (MS2): The structural fingerprint produced by collision-induced dissociation, used for metabolite identity confirmation
- Retention time (RT): The chromatographic elution position, which serves as an additional orthogonal identifier
This DDA phase functions as the "discovery engine" of the workflow, comprehensively cataloging the metabolic constituents present in the sample matrix without any prior assumptions about which compounds should be present.
Step 3: Metabolite Identification and Annotation
The raw DDA data — MS1 accurate masses, MS2 spectra, and retention times — are matched against a proprietary metabolite database using multi-dimensional scoring algorithms. Metabolite identification proceeds through tiered confidence levels based on Metabolomics Standards Initiative (MSI) guidelines (Sumner et al., 2007; Reisdorph et al., 2019):
- Level 1 (Confirmed identification): Matched by authentic chemical standard (exact RT + MS1 + MS2 alignment)
- Level 2 (Probable identification): Matched by MS1 accurate mass and MS2 spectral library similarity (without standard confirmation)
- Level 3 (Tentative candidate): Matched by MS1 accurate mass and putative annotation from databases or literature
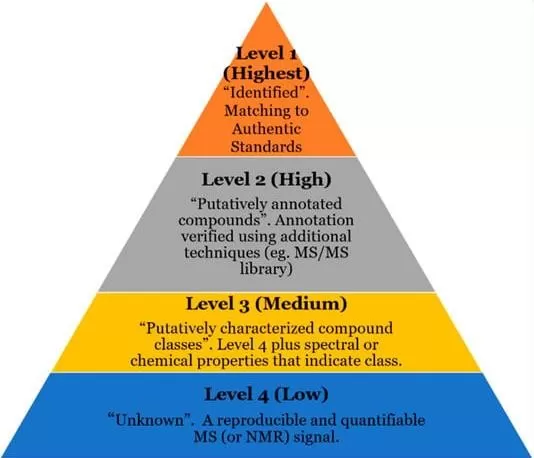
Figure 2. Currently accepted levels of confidence in metabolomics compound identification. Image reproduced from Reisdorph et al., 2019, Metabolites, 10(1), 8.
Once metabolites are annotated, the corresponding MRM transition parameters for those metabolites — including Q1/Q3 ion pairs, DP, and CE — can be retrieved from the database and carried forward into the quantitative workflow. In this way, the DDA phase not only determines which metabolites are present in the sample type, but also defines which database-supported targets will be monitored during MRM analysis.
Step 4: MRM-Based Metabolite Quantification on Triple Quadrupole MS
With the annotated metabolite list defined in the discovery phase, each individual study sample is then analyzed separately using the corresponding pre-optimized MRM transitions stored in the proprietary database. Samples undergo UPLC separation coupled to a triple quadrupole mass spectrometer (such as a QQQ or QTRAP instrument), which operates in multiple reaction monitoring (MRM) mode.
In MRM mode, the first quadrupole (Q1) selectively filters the specific precursor ion (Q1 m/z) for each target metabolite. The selected precursor then enters the collision cell, where it undergoes fragmentation under optimized collision energy (CE). The third quadrupole (Q3) subsequently monitors one or more characteristic product ions (Q3 m/z). This sequence of precursor ion selection, fragmentation, and product ion monitoring provides exceptional selectivity, effectively reducing matrix interference and background noise that often affect full-scan untargeted approaches (Tsakalof et al., 2024).
Because the MRM transitions monitored in this step were derived from actual metabolite identifications performed on the same sample type during the discovery phase, every monitored compound has already been confirmed to be genuinely present and detectable — unlike conventional targeted panels that may include irrelevant targets. This ensures that the MRM acquisition focuses exclusively on relevant, verifiable metabolites, maximizing both efficiency and data quality.
 mode_1776217801_WNo_710d429.webp)
Figure 3. Operation of triple quadrupole MS in Multiple Reaction Monitoring (MRM) mode. Image reproduced from Tsakalof et al., 2024, Molecules (Basel, Switzerland), 29(23), 5808.
Step 5: Quantitative Data Processing and Bioinformatics Analysis
Raw MRM signal data from all individual samples undergo peak integration, quality filtering, and normalization. Peak integration extracts the chromatographic peak area (or height) for each metabolite's MRM transition in each sample. Rigorous QC criteria are applied: metabolites showing poor peak shape, excessive variation in pooled QC samples (> 30% RSD), or detection rates below a threshold (e.g., < 50% of samples) are flagged or excluded.
Normalization corrects for systematic variations arising from sample preparation differences, injection volume fluctuations, and instrumental drift — commonly using QC-based robust LOESS (locally estimated scatterplot smoothing) signal correction or internal standard normalization. The final output is a quantitative metabolite matrix (samples × metabolites) suitable for downstream statistical analysis, including:
- Univariate statistics (t-test, ANOVA, fold-change analysis) to identify significantly altered metabolites
- Multivariate analysis (PCA, PLS-DA, OPLS-DA) to reveal metabolic clustering and biomarker patterns
- Pathway enrichment analysis (KEGG, HMDB, custom databases) to interpret biological meaning
- Integration with other omics datasets (transcriptomics, proteomics) for multi-omics systems biology insights
3. Advantages of Widely-Targeted Metabolomics Over Other Approaches
The value of widely-targeted metabolomics becomes clearer when it is compared with the two approaches it is most often positioned between: untargeted metabolomics and conventional targeted metabolomics. Rather than fully replacing either strategy, widely-targeted metabolomics occupies an intermediate analytical space. Its main strength lies in combining relatively broad metabolite coverage with a more controlled quantitative framework, making it particularly useful for studies that require both metabolic breadth and reproducible comparative analysis.
3.1 Widely-Targeted vs Untargeted Metabolomics
Untargeted metabolomics is well suited to open-ended discovery because it does not require a predefined metabolite list at the time of acquisition. This makes it especially valuable for exploratory studies, poorly characterized biological systems, and projects focused on unexpected metabolic features. However, its quantitative performance is often limited by full-scan acquisition, variable ionization efficiency, feature alignment challenges, and incomplete or inconsistent MS/MS coverage across large sample sets.
Widely-targeted metabolomics addresses some of these limitations by separating discovery from quantification. Metabolites are first detected and annotated by high-resolution DDA analysis, then measured across all samples by triple quadrupole MS in MRM mode. For the metabolites included in this workflow, this design generally improves signal consistency, reduces missing values, and supports more stable cross-sample comparison. At the same time, it remains less open than a fully untargeted strategy for compounds that are absent from the database or not captured during the discovery phase.
Untargeted vs Widely-Targeted Metabolomics: Key Analytical Differences
| Dimension | Untargeted Metabolomics | Widely-Targeted Metabolomics |
|---|---|---|
| Analytical goal | Broad, unbiased discovery | Broad profiling with targeted quantification |
| Acquisition mode | Full-scan MS, often with DDA/DIA MS/MS | DDA discovery followed by MRM measurement |
| Unknown metabolite capture | Stronger | More limited |
| Quantitative consistency | Moderate | Generally higher for monitored metabolites |
| Reproducibility across cohorts | Variable | Typically improved |
| Missing values | More common | Usually reduced |
| Low-abundance metabolite measurement | Often less stable | Often more sensitive |
| Best fit | Exploratory discovery | Comparative studies requiring breadth and consistency |
3.2 Widely-Targeted vs Targeted Metabolomics
Conventional targeted metabolomics is designed for accurate measurement of a fixed set of predefined metabolites, usually selected on the basis of pathway relevance, biological importance, or assay availability. Its principal advantage is analytical control: transitions are optimized in advance, performance can be validated in detail, and quantification is highly focused. For studies centered on a limited number of known metabolites, this remains one of the most robust analytical strategies.
Widely-targeted metabolomics differs from conventional targeted workflows in that its analyte panel is not defined entirely in advance. Instead, the metabolites selected for MRM quantification are informed by the discovery-phase results and matched to compounds supported by the database. This gives the method greater flexibility and broader chemical coverage than a fixed targeted panel. However, it should not be considered equivalent to fully validated targeted assays used for absolute quantification, regulatory testing, or clinical decision-making, where calibration models and isotope-labeled internal standards remain essential.
Targeted vs Widely-Targeted Metabolomics: Key Analytical Differences
| Dimension | Targeted Metabolomics | Widely-Targeted Metabolomics |
|---|---|---|
| Analytical goal | Precise measurement of predefined metabolites | Large-scale measurement of annotated metabolites |
| Panel design | Fixed in advance | Informed by discovery results |
| Metabolite coverage | Narrow to moderate | Broad |
| Flexibility across matrices | More limited | Generally higher |
| Unexpected metabolite inclusion | Limited | Greater, within database scope |
| Quantitative rigor | Highest in validated assays | Strong for large-scale relative quantification |
| Method expansion | Often labor-intensive | More scalable |
| Best fit | Focused hypothesis testing and validated panels | Broad profiling with quantitative comparability |
4. Applications of Widely-Targeted Metabolomics in Research
With its advantages in metabolite coverage, annotation confidence, and quantitative reproducibility, widely-targeted metabolomics has become a valuable tool in both biomedical and agricultural research. It is increasingly used in applications such as biomarker discovery, disease mechanism analysis, crop quality assessment, and stress-response studies, where large-scale yet reliable metabolite profiling is required.
4.1 Cancer Biomarker Discovery
Widely-targeted metabolomics is increasingly used in cancer biomarker research, especially when broad metabolic coverage and reproducible quantification are both required. In an open-access study on papillary thyroid carcinoma (PTC), plasma metabolomics and lipidomics were performed in 94 patients and 100 healthy controls. The analysis identified 113 differential metabolites and highlighted disruptions in branched-chain amino acid metabolism, glutamine metabolism, and the TCA cycle. Three metabolites — sebacic acid, L-glutamine, and indole-3-carboxaldehyde — showed promising diagnostic performance across discovery and validation cohorts (Wang et al., 2023). This case illustrates how widely-targeted metabolomics can support biomarker screening while preserving pathway-level interpretability.
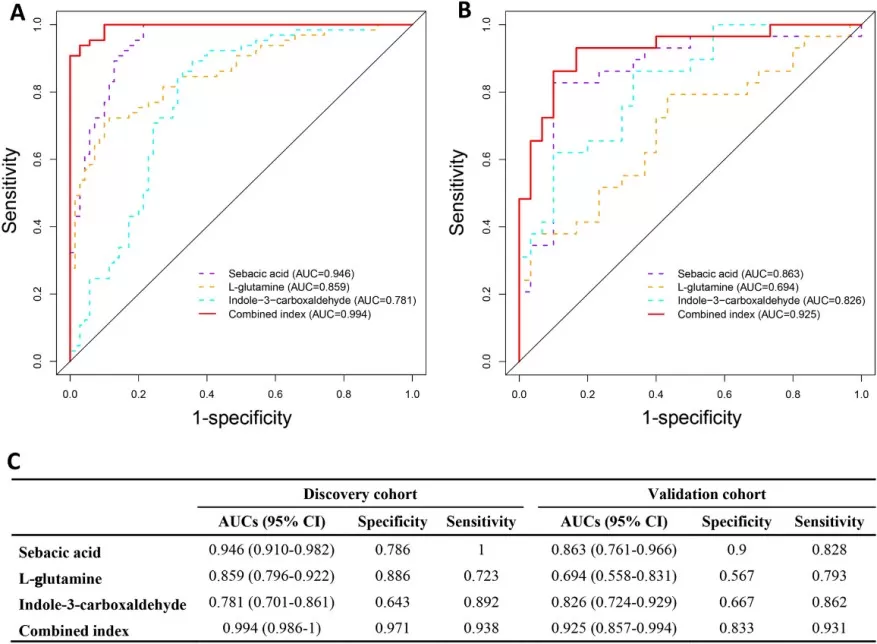
Figure 4. The ROC curves of the discriminative model showed three biomarkers — sebacic acid, L-glutamine, and indole-3-carboxaldehyde — and the combined indices in the discovery and validation cohorts. Image reproduced from Wang et al., 2023, Scientific Reports, 13(1), 17666.
4.2 Critical Care and Disease Stratification
Another important medical application is disease stratification in complex systemic disorders. A recent open-access study on sepsis-associated encephalopathy (SAE) applied widely targeted LC-MS/MS metabolomics to plasma from healthy controls, sepsis patients, and SAE patients. The workflow identified 12 discriminatory metabolites, with succinate showing a stepwise increase from health to sepsis to SAE and correlating with clinical severity. Follow-up experiments in a cecal ligation and puncture (CLP) mouse model further suggested a mechanistic link between succinate, neuroinflammation, and cognitive injury (Hu et al., 2025). This study shows that widely-targeted metabolomics can contribute not only to biomarker discovery, but also to biologically informed patient stratification in critical illness.
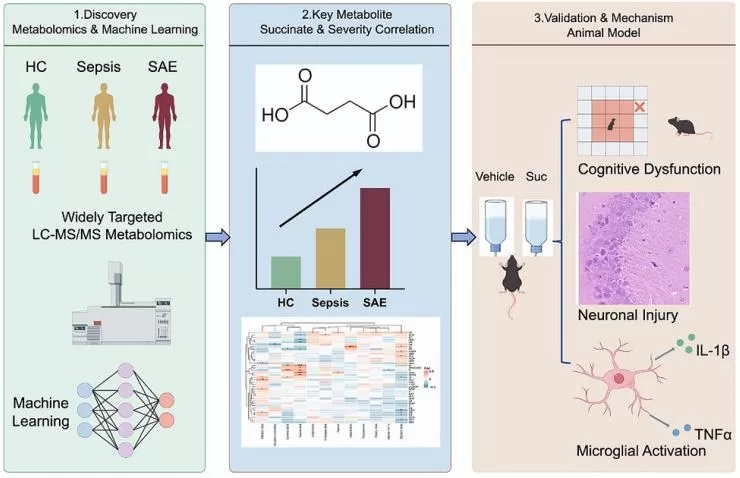
Figure 5. Succinate-Driven Metabolic Remodeling in Sepsis-Associated Encephalopathy. Image reproduced from Hu et al., 2025, iScience, 29(1), 114520.
4.3 Fruit Quality Evaluation in Horticultural Crops
In agronomy, widely-targeted metabolomics is well suited to dissecting quality traits in economically important crops. An open-access study on late-maturing hybrid mandarins compared fruits grafted onto four rootstocks using UPLC-MS/MS-based widely targeted metabolomics. The authors identified 1006 metabolites and found that rootstock choice influenced the accumulation of lipids, phenolic acids, and flavonoids, with pathway differences involving tryptophan metabolism, phenylpropanoid biosynthesis, and sphingolipid metabolism. These results provide a metabolite-level explanation for rootstock-dependent differences in fruit quality (Wang et al., 2024). The case illustrates the value of widely-targeted metabolomics for linking horticultural practices with nutritional and sensory traits.
4.4 Crop Stress Response and Resistance Mechanisms
Widely-targeted metabolomics is also useful for studying stress adaptation and resistance in agricultural systems. In an open-access study of Abutilon theophrasti exposed to glufosinate ammonium, researchers used widely targeted metabolomics to compare metabolic responses between two populations under the same herbicide treatment. A total of 2546 metabolites were detected, and key pathway shifts were observed in arginine and proline metabolism, amino acid biosynthesis, and D-amino acid metabolism. Rather than identifying a single resistance marker, the study mapped broader metabolic reprogramming associated with herbicide response (Guo et al., 2025). This example shows how widely-targeted metabolomics can help clarify stress-related mechanisms relevant to weed management and crop production systems.
MetwareBio: Your Trusted Partner for Widely-Targeted Metabolomics
As a global leader in metabolomics, proteomics, and multi-omics CRO services, MetwareBio provides end-to-end widely-targeted metabolomics solutions for both biomedical and plant research. Key advantages of our widely-targeted metabolomics services:
- Large in-house databases supporting broad metabolite coverage and confident annotation for both the Plant Widely-Targeted Metabolomics Service and the TM Widely-Targeted Metabolomics Service (for human/animal samples), with 61,000+ plant-associated metabolites and a dedicated biomedical widely-targeted metabolomics library.
- Complete widely-targeted metabolomics service workflow spanning project consultation, sample processing, LC-MS/MS analysis, quality control, bioinformatics, and biological interpretation.
- Expert technical team with deep experience in widely-targeted metabolomics method development, project execution, and data interpretation.
- Proven delivery record, with 20,000+ projects supported and 1,000+ publications enabled across diverse metabolomics and multi-omics applications.
Contact MetwareBio to discuss a widely-targeted metabolomics project or request a quote.
Contact UsReferences
- Chen, W., Gong, L., Guo, Z., Wang, W., Zhang, H., Liu, X., Yu, S., Xiong, L., & Luo, J. (2013). A novel integrated method for large-scale detection, identification, and quantification of widely targeted metabolites: application in the study of rice metabolomics. Molecular Plant, 6(6), 1769–1780. https://doi.org/10.1093/mp/sst080
- Reisdorph, N. A., Walmsley, S., & Reisdorph, R. (2019). A Perspective and Framework for Developing Sample Type Specific Databases for LC/MS-Based Clinical Metabolomics. Metabolites, 10(1), 8. https://doi.org/10.3390/metabo10010008
- Sumner, L. W., Amberg, A., Barrett, D., Beale, M. H., Beger, R., Daykin, C. A., Fan, T. W., Fiehn, O., Goodacre, R., Griffin, J. L., Hankemeier, T., Hardy, N., Harnly, J., Higashi, R., Kopka, J., Lane, A. N., Lindon, J. C., Marriott, P., Nicholls, A. W., Reily, M. D., … Viant, M. R. (2007). Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics: Official Journal of the Metabolomic Society, 3(3), 211–221. https://doi.org/10.1007/s11306-007-0082-2
- Tsakalof, A., Sysoev, A. A., Vyatkina, K. V., Eganov, A. A., Eroshchenko, N. N., Kiryushin, A. N., Adamov, A. Y., Danilova, E. Y., & Nosyrev, A. E. (2024). Current Role and Potential of Triple Quadrupole Mass Spectrometry in Biomedical Research and Clinical Applications. Molecules (Basel, Switzerland), 29(23), 5808. https://doi.org/10.3390/molecules29235808
- Wang, Z., Yang, Y., Xing, Y., Si, D., Wang, S., Lin, J., Li, C., Zhang, J., & Yin, D. (2023). Combined metabolomic and lipidomic analysis uncovers metabolic profile and biomarkers for papillary thyroid carcinoma. Scientific Reports, 13(1), 17666. https://doi.org/10.1038/s41598-023-41176-4
- Hu, H., Feng, Y., Zhou, Y., Peng, S., Li, D., Wu, S., Jiang, H., Lu, Y., Chen, J., Song, Y., & Zhu, W. (2025). Widely targeted metabolomics and machine learning identify succinate as a key metabolite in sepsis-associated encephalopathy. iScience, 29(1), 114520. https://doi.org/10.1016/j.isci.2025.114520
- Wang, M., Chen, Y., Li, S., Yu, J., Yang, L., & Hong, L. (2024). Widely Targeted Metabolomic Analysis Provides New Insights into the Effect of Rootstocks on Citrus Fruit Quality. Metabolites, 14(4), 242. https://doi.org/10.3390/metabo14040242
- Guo, X., Wang, Y., Guo, Y., Luo, C., & Cong, K. (2025). Widely Targeted Metabolomics Reveals Metabolic Divergence in Abutilon theophrasti Populations Under Glufosinate Ammonium Treatment. Plants (Basel, Switzerland), 14(13), 1994. https://doi.org/10.3390/plants14131994


