Plant metabolomics is one of the most informative approaches in modern plant biology. Plant metabolism spans a vast chemical space, encompassing an estimated 200,000 to 1,000,000 specialized metabolites and about 8,000 primary metabolites that shape plant growth, development, defense, and environmental adaptation. By profiling these molecules, plant metabolomics provides a direct biochemical readout of plant physiology and offers critical insights into the regulation of these processes. Recent advances in mass spectrometry-based workflows have transformed metabolite analysis from a relatively low-throughput activity into a scalable research strategy capable of detecting large numbers of features in a single experiment, driving the widespread adoption of plant metabolomics across plant research. This article provides a comprehensive guide to plant metabolomics, covering major metabolite classes, analytical platforms, research strategies, and real-world applications, aiming to equip researchers with the foundational knowledge needed to navigate this rapidly evolving field.
1. What Is Plant Metabolomics?
Plant metabolomics is the comprehensive qualitative and quantitative analysis of low-molecular-weight metabolites in plant cells, tissues, organs, or whole organisms. As a key branch of systems biology and an increasingly important component of functional genomics, it aims to characterize not only which metabolites are present, but also how their abundance and composition change across genotypes, developmental stages, and environmental conditions.
Because metabolites are closely linked to biochemical function, metabolomic data often provide one of the most direct molecular readouts of plant physiological state. Compared with upstream genomics and transcriptomics layers, metabolite profiles are highly dynamic and integrate both genetic regulation and environmental inputs, making plant metabolomics particularly valuable for studying growth, development, stress responses, quality traits, and metabolic regulation. In practical applications, plant metabolomics helps translate molecular variation into biologically interpretable biochemical change.
2. The Major Metabolite Classes in Plant Metabolism
As sessile organisms, plants have evolved extraordinary metabolic diversity as a chemical adaptation strategy to cope with environmental stress. This long evolutionary history has generated a vast plant metabolome: the number of plant specialized metabolites has been estimated to exceed 200,000, while the true biosynthetic capacity of the plant kingdom may reach 1,000,000 compounds (Huang & Dudareva, 2023).
To make this complexity more interpretable, plant metabolites are commonly grouped into primary metabolites and secondary metabolites, now more often referred to as specialized metabolites. Primary metabolites, such as sugars, amino acids, organic acids, and many lipids, are essential for energy metabolism, biosynthesis, growth, and development. Specialized metabolites, in contrast, are more closely associated with defense, chemical communication, and environmental adaptation (Huang & Dudareva, 2023; Hao et al., 2025). Within this primary–specialized framework, major metabolite classes define much of the biochemical landscape explored in plant metabolomics studies:
Major Metabolite Classes in Plant Metabolism and Their Biological Functions
| Major metabolite class | Representative compounds | Main physiological or biological functions |
|---|---|---|
| Carbohydrates | Glucose, fructose, sucrose | Energy supply, carbon skeleton provision, osmotic regulation, biomass accumulation |
| Amino acids | Glutamate, aspartate, proline, GABA | Protein synthesis, nitrogen metabolism, stress signaling, metabolic regulation |
| Lipids | Fatty acids, glycerolipids, phospholipids, sterols | Membrane structure, energy storage, signaling, cellular organization |
| Phenolics | Phenylpropanoids, lignin, flavonoids, anthocyanins, tannins | UV protection, antioxidant defense, pigmentation, pathogen defense, structural support |
| Terpenoids | Monoterpenes, sesquiterpenes, diterpenes, triterpenes, carotenoids | Defense against herbivores and pathogens, allelopathy, pollinator attraction, photoprotection, signaling |
| Alkaloids | Nicotine, caffeine, quinine, vinblastine | Chemical defense, allelopathic interaction, antimicrobial activity, ecological adaptation |
| Plant hormones | Auxin, gibberellins, cytokinins, abscisic acid, jasmonates, salicylic acid | Regulation of growth and development, stress signaling, coordination of physiological responses |
| Volatile organic compounds (VOCs) | Isoprene, monoterpenes, green leaf volatiles | Plant–plant communication, indirect defense, pollinator attraction, plant–microbe interaction |
3. Analytical Platforms for Plant Metabolomics
The chemical diversity of plant metabolites makes platform selection a central issue in plant metabolomics. Because plant compounds differ widely in polarity, volatility, thermal stability, and abundance, no single analytical platform can capture the full plant metabolome. As a result, modern plant metabolomics relies on complementary analytical platforms, among which liquid chromatography-mass spectrometry (LC-MS), gas chromatography-mass spectrometry (GC-MS), and nuclear magnetic resonance (NMR) spectroscopy remain the most widely used for metabolite profiling, compound annotation, and biological interpretation (Hao et al., 2025; Patel et al., 2021). (Learn more at: LC-MS VS GC-MS: What's the Difference)
3.1 LC-MS/MS: The Most Versatile Platform for Plant Metabolomics
Liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS) is the most widely used platform in plant metabolomics because of its high sensitivity and broad metabolite coverage. Coupled with electrospray ionization (ESI), LC-MS can analyze metabolites ranging from polar small molecules to non-polar lipids without derivatization, making it suitable for both untargeted metabolomics profiling and targeted quantification of flavonoids, alkaloids, phenolic acids, saponins, and phytohormones. High-resolution mass spectrometers such as Orbitrap and Q-TOF systems are widely used in untargeted metabolomics, offering high sensitivity, selectivity, and exceptional mass accuracy (Maisl et al., 2025). Ultra-performance liquid chromatography (UPLC) further improves separation efficiency by using sub-2 µm particles, enabling higher resolution and faster analysis than conventional HPLC systems. The integration of LC-MS with ion mobility spectrometry (IMS) has recently emerged as an advanced analytical approach that provides an additional dimension of separation for metabolites and their isomers, helping address the long-standing challenge of distinguishing structurally similar compounds that co-elute during chromatographic separation (Luo et al., 2023). Key limitations of LC-MS include matrix effects, incomplete spectral libraries, and limited suitability for highly volatile compounds.
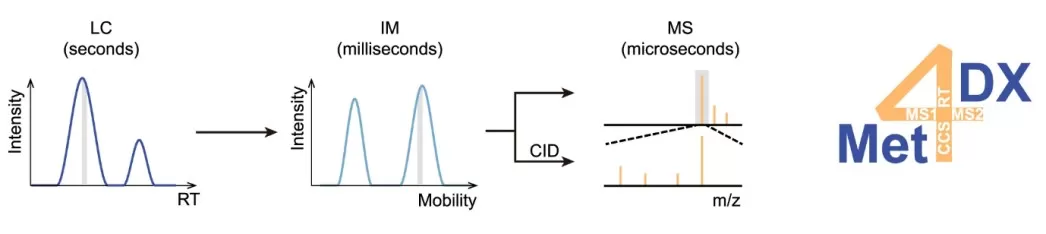
Figure 1. The multidimensional separations and measurements of metabolites using LC–IM–MS/MS technology. Image reproduced from Luo et al., 2023, Nature communications, 14(1), 1813.
3.2 GC-MS: A Core Platform for VOCs and Primary Metabolites
Gas chromatography-mass spectrometry (GC-MS) represents the most mature and well-established analytical platform in metabolomics, offering unparalleled chromatographic resolution and reproducible fragmentation patterns. GC-MS is particularly suited for the analysis of volatile organic compounds (VOCs) and low-molecular-weight primary metabolites, including amino acids, organic acids, sugars, and fatty acids. The technique achieves separation based on analyte volatility and interaction with the stationary phase, followed by electron ionization (EI) that produces highly characteristic and reproducible fragmentation spectra. This standardized ionization approach has enabled the development of extensive commercial and public spectral libraries—such as the NIST and Golm Metabolome Database—that facilitate confident metabolite identification through spectral matching (Frolova et al., 2025). The primary limitation of GC-MS lies in its restricted analyte scope: non-volatile and thermally labile metabolites require chemical derivatization prior to analysis, a process that adds sample preparation time, introduces potential artifacts, and may obscure the detection of certain compound classes. Consequently, GC-MS is most effectively deployed as a complementary technique within multi-platform workflows rather than as a standalone solution for comprehensive metabolome coverage.
3.3 NMR Spectroscopy: The Quantitative and Structural Complement
Nuclear magnetic resonance (NMR) spectroscopy occupies a unique position in the plant metabolomics toolkit, distinguished by its non-destructive nature, exceptional quantitative accuracy, and unparalleled capacity for de novo structure elucidation. Unlike mass spectrometry-based techniques, NMR does not require chromatographic separation or analyte ionization, thereby eliminating matrix effects and enabling direct, absolute quantification of metabolites without the need for authentic standards. NMR's ability to provide detailed chemical insights, including atom connectivity and stereochemical information, is crucial in obtaining comprehensive metabolite profiles and resolving structural ambiguities that MS alone cannot address (Kumar et al., 2026). Despite these strengths, NMR remains constrained by lower sensitivity, more limited metabolome coverage, substantial signal overlap in complex mixtures, and greater demands on instrument performance, sample amount, and acquisition time than MS-based metabolomics.
LC-MS vs. GC-MS vs. NMR: Strengths and Limitations
| Platform | Strengths | Limitations | Typical plant applications |
|---|---|---|---|
| LC-MS | High sensitivity, broad coverage, MS/MS-based annotation | Complex data processing, matrix effects, many features remain putative | Flavonoids, alkaloids, phenolics, lipids, semi-polar primary metabolites |
| GC-MS | Excellent reproducibility, strong libraries for small derivatized analytes and volatiles | Requires volatility or derivatization, narrower chemical coverage | Organic acids, amino acids, sugars, VOCs |
| NMR | Highly reproducible, structurally informative, quantitatively robust when standardized | Lower sensitivity than MS-based methods | Targeted quantification, metabolite confirmation, food and quality studies |
4. Research Strategies in Plant Metabolomics
Plant metabolomics is not defined by a single experimental design. Instead, researchers choose among multiple analytical strategies according to research goals. Targeted, untargeted, and widely targeted approaches differ fundamentally in their scope, quantification capabilities, and discovery potential. More recently, advances in spatial resolution and single-cell analysis have enabled the interrogation of metabolic heterogeneity across tissues and cell types, while multi-omics integration frameworks provide a systems-level view connecting metabolites to their genetic and regulatory underpinnings.
4.1 Targeted, Untargeted, and Widely Targeted Metabolomics
Targeted metabolomics quantifies predefined metabolites with high accuracy and sensitivity using MRM and authentic standards, ideal for hypothesis validation but limited to known compounds. Untargeted metabolomics detects thousands of features without prior knowledge, enabling discovery but often lacking confident identification and precise quantification. Widely targeted metabolomics bridges these approaches, combining the breadth of untargeted acquisition with calibration-based quantification for hundreds of metabolites. This hybrid strategy offers high throughput, broad coverage, and reliable quantitation, making it particularly valuable for systematic profiling of plant specialized metabolites and biomarker discovery.
Comparison of Targeted, Untargeted, and Widely Targeted Metabolomics
| Strategy | Detection scope | Quantitative profile | Primary use case |
|---|---|---|---|
| Untargeted | Broad survey of detectable features | Mostly relative | Hypothesis generation, biomarker discovery |
| Targeted | Predefined metabolite panel | High quantitative reliability | Validation, mechanistic testing, focused quantification |
| Widely-targeted | Large annotated metabolite panels | High-quality relative quantification | Large-cohort plant profiling and pathway-oriented screening |
4.2 Single-Cell Metabolomics: Resolving Cellular Heterogeneity
Bulk tissue analysis often obscures the strong cell-type specificity of plant metabolism, especially for specialized metabolites. Single-cell metabolomics addresses this limitation by measuring metabolites directly at the level of individual cells, making it possible to resolve metabolic heterogeneity that is otherwise lost after tissue homogenization. A recent advance came from a 2025 PNAS study that multiplexed single-cell metabolome and RNA-seq measurements in the same plant cells and used Catharanthus roseus as a model, providing a direct framework for linking transcriptional states with cell-specific metabolite accumulation (Kang et al., 2025).
4.3 Spatial Metabolomics and Mass Spectrometry Imaging
While single-cell approaches resolve cellular heterogeneity, spatial metabolomics preserves tissue context by visualizing metabolites in situ. In plant research, this strategy is largely driven by mass spectrometry imaging (MSI) platforms such as MALDI-MSI and related ambient or substrate-assisted imaging methods, which reveal where metabolites accumulate across tissues, organs, or living plant surfaces. Recent plant-focused reviews and methodological studies show that spatial metabolomics is expanding rapidly because it can uncover tissue-specific metabolic organization and can now be extended to fragile, water-rich, or difficult-to-slice plant materials through flexible-substrate imaging platforms (Li et al., 2025; Zhang et al., 2025).
4.4 Multi-Omics Integration for Systems-Level Interpretation
No single omics layer can fully explain plant phenotype. For this reason, metabolomics is increasingly integrated with genomics, transcriptomics, and proteomics to reconstruct regulatory links from gene variation to metabolic output. In plants, mGWAS and mQTL analyses have become important frameworks for associating metabolite variation with biosynthetic genes and regulatory loci, while newer computational approaches are using artificial intelligence and machine learning to improve metabolite annotation, pathway prediction, and candidate gene discovery. These integrative strategies are moving plant metabolomics from descriptive profiling toward systems-level biological interpretation (Zhu et al., 2023; Chi et al., 2024; Bai et al., 2024).
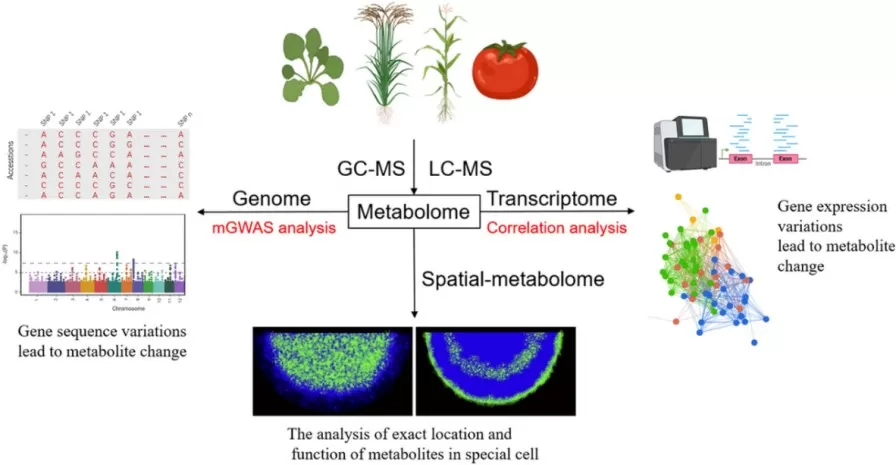
Figure 2. Pipeline of the integrated analysis of metabolome/genome/transcriptome to identify the primary and secondary pathways and genes. Image reproduced from Zhu et al., 2023, aBIOTECH, 4(1), 47–56.
5. Plant Metabolomics Workflow: From Sample to Insight
A robust metabolomics workflow is essential for converting plant samples into biologically meaningful results. Although protocols vary by platform and study design, most plant metabolomics studies follow a common sequence that includes experimental design, sample collection, sample preparation, metabolite extraction, data acquisition, data processing, metabolite annotation, and statistical analysis (Hao et al., 2025). Each step influences metabolome coverage, data quality, and the reliability of downstream biological interpretation.
Step 1: Experimental Design and Sample Collection
The workflow begins with a clearly defined biological question, which determines whether the study is discovery-driven or validation-oriented and guides the choice of sample type, controls, and biological replicate number. For plant samples, metabolism must be quenched rapidly at harvest, typically by immediate liquid-nitrogen freezing followed by storage at −80 °C, to minimize post-harvest metabolic turnover and degradation.
Step 2: Sample Preparation and Metabolite Extraction
After collection, tissues are ground and homogenized under cold conditions, and extraction solvents are selected according to metabolite polarity, commonly using methanol-, water-, or chloroform-based systems. For GC-MS, derivatization is usually required before analysis. In plant samples, effective cell-wall disruption and control of metabolite degradation are critical because extraction efficiency strongly shapes the detectable metabolome.
Step 3: Data Acquisition
The analytical platform is then selected according to the research objective: LC-MS for broad coverage of semi-polar and non-volatile metabolites, GC-MS for volatile and primary metabolites, and NMR for robust quantification and structural support. Instrument parameters and pooled quality-control samples are used to monitor analytical stability and batch consistency throughout acquisition.
Step 4: Data Processing and Metabolite Annotation
Raw data are converted into interpretable feature tables through peak detection, alignment, filtering, and normalization. Metabolite annotation then relies on public and commercial libraries, including METLIN, MassBank, and GNPS, as well as software such as XCMS, MZmine, and MetaboAnalyst. A major bottleneck remains that many detected features cannot yet be confidently matched to known metabolites.
Step 5: Statistical Analysis and Biological Interpretation
The final stage uses statistical analysis to extract biological meaning from the processed data. Common approaches include PCA, PLS-DA, OPLS-DA, differential metabolite screening, and pathway enrichment, with the ultimate goal of identifying phenotype-associated metabolic markers or regulatory networks.
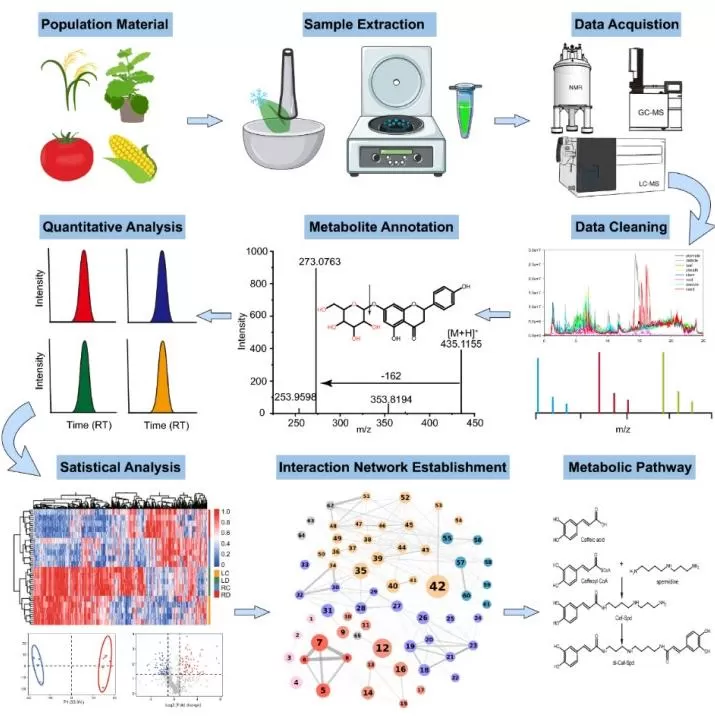
Figure 3. Diagram of plant metabolomics workflow. Image reproduced from Hao et al., 2025, aBIOTECH, 6(1), 116–132.
6. Applications of Plant Metabolomics in Research and Agriculture
Metabolomics has transitioned from an emerging technology to an indispensable tool for addressing fundamental and applied questions in plant biology. Its capacity to capture the biochemical phenotype of plants under diverse conditions enables applications spanning from basic stress physiology to translational crop improvement, natural product discovery, and rhizosphere ecology.
6.1 Stress Response and Environmental Adaptation
In stress biology, plant metabolomics is widely used to reveal how plants reprogram metabolism under drought, salinity, heat, cold, or heavy-metal stress, making it valuable for identifying metabolic markers of tolerance and clarifying adaptive pathways. In a salt-stress study of perennial ryegrass, Cao et al. (2024) combined metabolomics with transcriptomics and showed that the salt-tolerant genotype accumulated higher levels of phenylpropanoids, flavonoids, and anthocyanins and displayed stronger activation of the corresponding biosynthetic genes than the salt-sensitive genotype. The study identified phenylpropanoid biosynthesis as a major pathway underlying salt adaptation in this species.
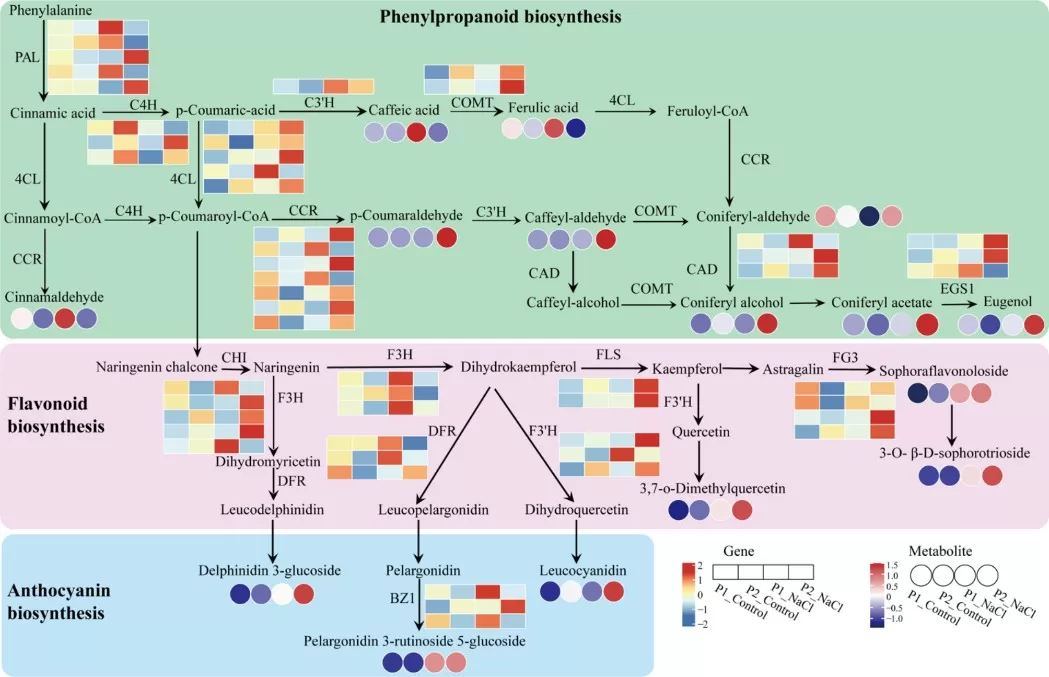
Figure 4. Phenylpropanoid, flavonoid and anthocyanin biosynthesis network diagram in salt sensitive genotype P1 and salt tolerant genotype P2 of perennial ryegrass. Image reproduced from Cao et al., 2024, BMC plant biology, 24(1), 1225.
6.2 Crop Improvement and Quality Breeding
In crop breeding, plant metabolomics supports high-throughput phenotyping for flavor, nutrition, and stress-related traits, and helps translate biochemical variation into practical breeding targets. A representative example is the citrus fruit-quality study by Wang et al. (2024), which used widely targeted metabolomics to analyze three late-maturing mandarin varieties grafted onto four rootstocks. The authors identified 1,006 metabolites and showed that rootstock-dependent differences were concentrated in lipids, phenolic acids, and flavonoids, demonstrating how metabolomics can reveal biochemical pathways relevant to fruit quality and guide rootstock selection in quality breeding.
6.3 Natural Product Discovery in Medicinal Plants
For natural product discovery, metabolomics enables efficient chemical mapping of medicinal plants and accelerates the identification of potentially bioactive compounds. In an AI-guided LC-MS/MS study of three medicinal Helichrysum species, Lephatsi et al. (2025) used molecular networking and complementary extraction strategies to expand metabolome coverage and showed that the metabolomes were rich in phenylpropanoids, lipids, and lipid-like molecules, while also differing in flavonoid glucoside and triterpenoid profiles across species. This work demonstrates how medicinal plant metabolomics can uncover species-specific chemical fingerprints and broaden the discovery space for pharmacologically relevant natural products.
6.4 Plant–Microbe Interactions and Rhizosphere Metabolism
In plant–microbe interaction research, rhizosphere metabolomics is used to trace how root exudate chemistry shapes belowground ecological interactions and disease outcomes. A recent rice intercropping study by Han et al. (2025) showed that varietal intercropping altered root exudate composition, enhanced blast resistance, and identified core metabolites—including azelaic acid, suberic acid, betaine, and phenyl acetate—that were positively associated with immune-related gene expression. Functional validation further showed that these metabolites inhibited Magnaporthe oryzae growth or reduced rice blast incidence after exogenous application, providing direct evidence that root-exuded metabolites can mediate belowground disease resistance.
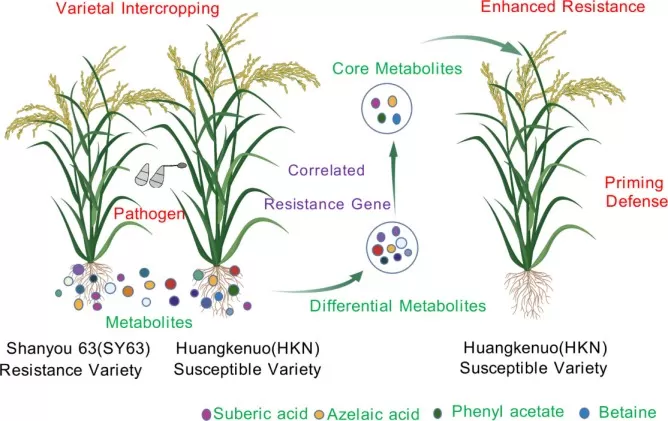
Figure 5. The schematic diagram of metabolite-driven varietal intercropping-induced resistance to rice blast. Image reproduced from Han et al., 2025, BMC plant biology, 25(1), 1730.
7. Challenges and Future Directions in Plant Metabolomics
Despite rapid technical progress, several core metabolomics challenges still limit the full potential of plant metabolomics. The most fundamental bottleneck is incomplete reference knowledge: current analytical platforms can detect thousands of metabolic features from a single sample, but only a minority can be confidently annotated because plant chemical diversity far exceeds the coverage of existing spectral libraries, plant-specific databases, and authentic standards. In practice, unknown metabolite identification remains the central obstacle linking limited metabolome coverage, low annotation rates, and reduced biological interpretability. A second major challenge is data integration and standardization. Differences in extraction protocols, instrument settings, annotation criteria, and reporting practices still reduce comparability and reproducibility across platforms, laboratories, and studies.
Looking ahead, the future of plant metabolomics will likely be shaped by more routine single-cell and spatial multi-omics, broader comparative plant metabolome resources, and more standardized data-sharing frameworks. At the same time, AI in metabolomics is expected to accelerate automated metabolite annotation, retention time prediction, spectral interpretation, and structural prediction for unknown compounds, helping move plant metabolomics from large-scale feature detection toward deeper functional interpretation.
Advance Your Plant Metabolomics Research with MetwareBio
Plant metabolomics has become an indispensable tool for linking plant biochemistry with physiology, stress adaptation, quality traits, and natural product discovery. MetwareBio is a global leader in metabolomics services and multi-omics solutions, dedicated to advancing plant and life science research through innovative technologies.
MetwareBio's Metabolomics Services and Multi-Omics solutions provide comprehensive support for your plant research. Our key strengths include:
- A proprietary plant metabolite database covering 61,000+ plant-associated metabolites
- Patented widely-targeted metabolomics for broad coverage and reliable quantification
- Comprehensive platforms, including LC-MS, GC-MS, spatial metabolomics, targeted, untargeted, and widely targeted metabolomics for plants
- Integrated multi-omics capability with transcriptomics, proteomics, microbiome, and genomics workflows
- Proven experience, with 20,000+ projects supported and 1,000+ publications enabled
If you are interested in plant metabolomics or multi-omics integration research, please do not hesitate to contact us.
Contact UsReferences
- Hao, Y., Zhang, Z., Luo, E., Yang, J., & Wang, S. (2025). Plant metabolomics: applications and challenges in the era of multi-omics big data. aBIOTECH, 6(1), 116–132. https://doi.org/10.1007/s42994-024-00194-0
- Huang, X. Q. & Dudareva, N. (2023). Plant specialized metabolism. Current Biology, 33(11), R473–R478. https://doi.org/10.1016/j.cub.2023.01.057
- Patel, M. K., Pandey, S., Kumar, M., Haque, M. I., Pal, S., & Yadav, N. S. (2021). Plants metabolome study: emerging tools and techniques. Plants (Basel, Switzerland), 10(11), 2409. https://doi.org/10.3390/plants10112409
- Luo, M., Yin, Y., Zhou, Z., Zhang, H., Chen, X., Wang, H., & Zhu, Z. J. (2023). A mass spectrum-oriented computational method for ion mobility-resolved untargeted metabolomics. Nature communications, 14(1), 1813. https://doi.org/10.1038/s41467-023-37539-0
- Maisl, C., Schuhmacher, R., & Bueschl, C. (2025). Accuracy, linearity, and statistical differences in comparative quantification in untargeted plant metabolomics using LC-ESI-Orbitrap-MS. Analytical and bioanalytical chemistry, 417(11), 2293–2309. https://doi.org/10.1007/s00216-025-05818-y
- Frolova, N., Orlova, A., Popova, V., Bilova, T., & Frolov, A. (2025). Gas chromatography-mass spectrometry (GC-MS) in the plant metabolomics toolbox: sample preparation and instrumental analysis. Biomolecules, 16(1), 16. https://doi.org/10.3390/biom16010016
- Kumar, N., & Jaitak, V. (2026). Recent advancement in NMR based plant metabolomics: techniques, tools, and analytical approaches. Critical reviews in analytical chemistry, 56(1), 1–25. https://doi.org/10.1080/10408347.2024.2375314
- Kang, M., Vu, A. H., Casper, A. L., Kim, R., Wurlitzer, J., Heinicke, S., Yeroslaviz, A., Caputi, L., & O'Connor, S. E. (2025). Single-cell metabolome and RNA-seq multiplexing on single plant cells. Proceedings of the National Academy of Sciences of the United States of America, 122(43), e2512828122. https://doi.org/10.1073/pnas.2512828122
- Zhang, G., Zheng, H., Wang, X., Han, S., Liu, W., Sun, C., Hu, Q., & Ma, C. (2025). Flexible substrate-based mass spectrometry platform for in situ non-destructive molecular imaging of living plants. Plant biotechnology journal, 23(1), 97–111. https://doi.org/10.1111/pbi.14482
- Li, R., Wang, F., & Wang, J. (2025). Spatial metabolomics and its application in plant research. International journal of molecular sciences, 26(7), 3043. https://doi.org/10.3390/ijms26073043
- Zhu, F., Wen, W., Cheng, Y., Alseekh, S., & Fernie, A. R. (2023). Integrating multiomics data accelerates elucidation of plant primary and secondary metabolic pathways. aBIOTECH, 4(1), 47–56. https://doi.org/10.1007/s42994-022-00091-4
- Chi, J., Shu, J., Li, M., Mudappathi, R., Jin, Y., Lewis, F., Boon, A., Qin, X., Liu, L., & Gu, H. (2024). Artificial intelligence in metabolomics: a current review. Trends in analytical chemistry : TRAC, 178, 117852. https://doi.org/10.1016/j.trac.2024.117852
- Bai, W., Li, C., Li, W., Wang, H., Han, X., Wang, P., & Wang, L. (2024). Machine learning assists prediction of genes responsible for plant specialized metabolite biosynthesis by integrating multi-omics data. BMC genomics, 25(1), 418. https://doi.org/10.1186/s12864-024-10258-6
- Cao, Y. H., Lü, Z. L., Li, Y. H., Jiang, Y., & Zhang, J. L. (2024). Integrated metabolomic and transcriptomic analysis reveals the role of root phenylpropanoid biosynthesis pathway in the salt tolerance of perennial ryegrass. BMC plant biology, 24(1), 1225. https://doi.org/10.1186/s12870-024-05961-1
- Wang, M., Chen, Y., Li, S., Yu, J., Yang, L., & Hong, L. (2024). Widely targeted metabolomic analysis provides new insights into the effect of rootstocks on citrus fruit quality. Metabolites, 14(4), 242. https://doi.org/10.3390/metabo14040242
- Lephatsi, M. M., Choene, M. S., Kappo, A. P., Madala, N. E., & Tugizimana, F. (2025). Mapping the Helichrysum metabolome: uncovering species-specific chemistry through an AI-guided LC-MS/MS workflow. Molecular omics, 21(6), 747–759. https://doi.org/10.1039/d5mo00118h
- Han, G. Y., Li, R. P., Zhu, X. Q., Hendriks, P. W., Tang, M., Li, B., Xie, Y., Zan, Q. A., Wang, Y. Y., & He, P. (2025). Rice varietal intercropping mediates resistance to rice blast (Magnaporthe oryzae) through core root exudates. BMC plant biology, 25(1), 1730. https://doi.org/10.1186/s12870-025-07724-y


