Multi-omics correlation analysis has become an important strategy for connecting transcriptomics, proteomics, and metabolomics in the same biological system. Single-omics studies can reveal valuable molecular changes, but they rarely explain how signals propagate across layers or where regulation becomes decoupled. This is especially important in cancer, metabolic disease, immunology, and crop science, where phenotypes often emerge from coordinated changes rather than from one molecular layer alone. A well-designed multi-omics correlation workflow can help identify shared molecular programs, prioritize biomarker candidates, and generate more testable biological hypotheses. This article provides a practical guide to multi-omics correlation analysis, covering core principles, method frameworks, application-driven strategy selection, and critical workflow factors that shape analytical robustness and biological interpretability.
1. What Is Multi-Omics Correlation Analysis?
Multi-omics correlation analysis is more than a generic data-integration step. In practical research, multi-omics correlation analysis refers to a group of computational strategies used to detect coordinated variation across matched omics layers such as genes, proteins, metabolites, lipids, or epigenetic features measured from the same biological system and to translate those relationships into biologically meaningful structure. The central goal is not simply to stack datasets together, but to determine how signals observed in one molecular layer are reflected, modified, or buffered in another.
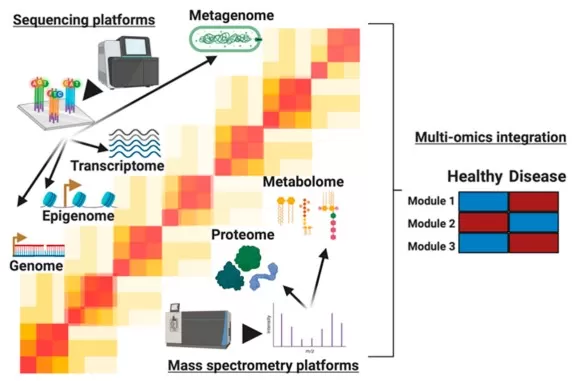
Figure 1. The molecules profiled in multi-omics studies. Image adapted from Lancaster et al. (2020), Biomolecules, 10(12), 1606.
That distinction is important because mRNA abundance, protein abundance, and metabolite levels are related but not interchangeable (Liu et al., 2016). Transcript levels often correlate only modestly with protein abundance, while metabolite levels can depend on enzyme activity, substrate availability, transport, compartmentalization, and pathway context rather than on one upstream molecule alone.
2. Core Method Families for Multi-Omics Correlation Analysis
Based on the type of structure being modeled, multi-omics correlation methods can be grouped into four main families. Each family captures a different layer of cross-omics organization and is best suited to a different research goal.
2.1 Feature-Level Correlation Methods: Pearson and Spearman Correlation
Feature-level association methods are the most direct form of multi-omics correlation analysis. They test whether two variables co-vary across matched samples, usually with Pearson correlation, Spearman correlation, robust correlation, or partial correlation. These methods are widely used in early-stage exploratory analysis because they are simple, intuitive, and easy to visualize as heatmaps, pairwise matrices, circos plots, or filtered edge lists. They are especially useful when the biological question centers on specific molecular relationships, such as transcript–protein concordance or associations between enzymes and metabolites.
Among these methods, Pearson or Spearman correlation remains the most common starting point. Pearson correlation is useful when the relationship is approximately linear and the data have been properly normalized and transformed, whereas Spearman correlation is often preferred for monotonic associations, especially when nonnormality, ordinal structure, or outliers are concerns (Schober et al., 2018; de Winter et al., 2016). In multi-omics studies, this layer is valuable for generating interpretable candidate pairs, but it also has clear limitations: feature-by-feature testing scales poorly in high dimensions, indirect associations are common, and many biologically important relationships are many-to-many rather than one-to-one. For that reason, feature-level analysis is best treated as an entry point rather than a complete integration strategy.
Other feature-level methods, such as partial correlation or conditional association analysis, can help reduce confounding from shared drivers, but they generally require stronger sample size support and more careful model assumptions. These methods are useful extensions, but they are less often the main framework in published multi-omics studies than simple pairwise correlation or more structured multivariate integration.
2.2 Latent-Variable Integration Methods: Why Sparse CCA Matters Most
Latent-variable integration methods are designed for a different question: instead of asking whether one feature correlates with another, they ask whether multiple omics layers share a smaller number of underlying variation patterns. This is often the most informative strategy when the number of variables greatly exceeds the number of samples, or when cross-omics structure is expected to be distributed across many features rather than concentrated in one pair. This family includes CCA, sparse CCA, PLS-related methods, O2PLS, and phenotype-guided frameworks such as DIABLO (Jiang et al., 2023; Bouhaddani et al., 2016; Singh et al., 2019).
Within this family, sparse canonical correlation analysis (sCCA) is a particularly important representative because it captures the core logic of multi-omics latent integration. Classical CCA identifies linear combinations of variables from two datasets that are maximally correlated. In omics studies, however, classical CCA is often unstable because the number of variables is very large relative to the number of samples. Sparse CCA addresses this problem by adding sparsity constraints so that only a subset of features contributes to each canonical component. This improves interpretability and helps identify cross-omics feature sets associated with the same biological axis. For studies focused on shared transcriptome–proteome or transcriptome–metabolome structure, sCCA is often one of the most principled unsupervised starting points (Witten & Tibshirani, 2009; Jiang et al., 2023).
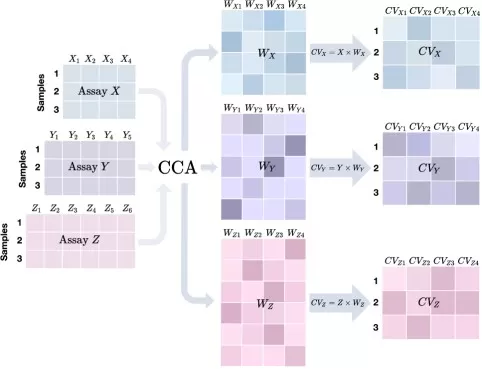
Figure 2. Cartoon illustration of a typical CCA-based method for three assays. Image adapted from Jiang et al. (2023), PLoS Genetics, 19(5), e1010517.
Other methods in this family are useful when the objective changes. O2PLS is valuable when one wants to separate shared structure from orthogonal, dataset-specific variation; this can be especially helpful when integrating datasets with different noise structures or very different assay behavior. DIABLO adds supervision by identifying cross-omics components that both capture shared information and discriminate between predefined phenotypic groups, making it well suited for multi-omics biomarker discovery and classification tasks. In other words, sCCA is a strong choice for discovering shared cross-omics axes, O2PLS is helpful when shared and private variation must be disentangled, and DIABLO becomes attractive when phenotype separation is part of the study goal.
2.3 Factor-Model Decomposition: MOFA+ for Shared and Private Signals
Factor-model approaches go one step further by explicitly decomposing each dataset into shared factors and omics-specific factors. Instead of maximizing pairwise correlation directly, these methods ask how much of the observed variation is common across datasets and how much is private to one data layer. This framing is especially helpful in real biological studies, where not all variation should be expected to align across transcriptomics, proteomics, and metabolomics (Brown et al., 2023; Argelaguet et al., 2020).
A useful representative here is MOFA+. Although its best-known application is in multi-modal single-cell analysis, the conceptual framework is broader: it learns latent factors that explain variation across multiple views, supports missing data patterns, and helps distinguish globally shared signals from view-specific structure (Argelaguet et al., 2020). For studies that aim to characterize system architecture rather than only rank cross-omics pairs, factor-model approaches can be more informative than simple correlation matrices. Recent alternatives such as MCFA further extend this logic in population-scale multi-omics settings and provide an interpretable way to separate correlated and private structure across many datasets (Brown et al., 2023).
 provides an unsupervised framework for the integration of multi-group and multi-view single-cell data_1778124169_WNo_871d666.webp)
Figure 3. Multi-Omics Factor Analysis v2 (MOFA+) provides an unsupervised framework for the integration of multi-group and multi-view single-cell data. Image adapted from Argelaguet et al. (2020), Genome Biology, 21(1), 111.
2.4 Network and Module Analysis: WGCNA for Cross-Omics Module Discovery
Network and module methods are designed for questions at the system level. Rather than prioritizing individual correlations or latent axes alone, they seek groups of molecules that behave coordinately and may act as regulatory or functional units. These approaches are especially attractive when the desired output is a module, hub, or pathway-oriented interpretation rather than a small list of molecular pairs (Langfelder & Horvath, 2008).
Weighted Gene Co-expression Network Analysis (WGCNA) remains the most established workhorse in this family. WGCNA constructs weighted correlation networks, clusters features into modules, summarizes each module with an eigengene, and then relates those modules to traits or external data. In multi-omics settings, this can be extended by building modules within each omics layer and then correlating module summaries across layers, or by using module logic to organize integrated interpretation. WGCNA is widely used because it is intuitive, scalable, and biologically interpretable. Its main caveat is sensitivity to preprocessing, sample size, and correlation structure. A visually dense network is not necessarily a stable one, so WGCNA results should always be combined with resampling, biological review, and conservative thresholding (Langfelder & Horvath, 2008).
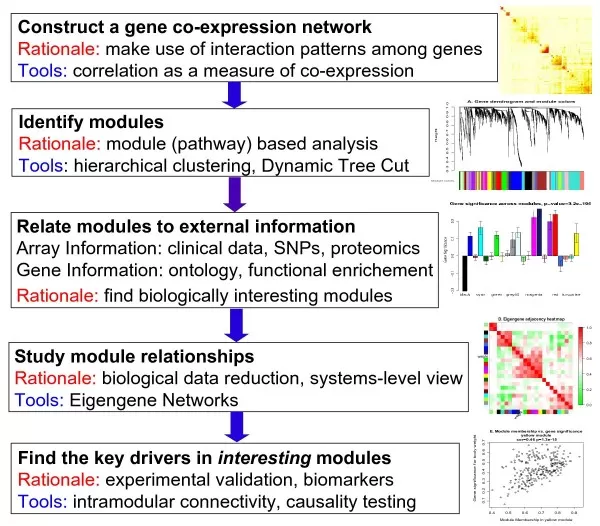
Figure 4. Overview of WGCNA methodology. Image adapted from Langfelder & Horvath (2008), BMC Bioinformatics, 9, 559.
3. How to Choose the Right Multi-Omics Correlation Analysis Method
Choosing the right method for multi-omics correlation analysis is rarely a matter of selecting the most advanced algorithm. Different method families are designed to capture different types of cross-omics structure, and each produces a different kind of biological output. For this reason, method selection is usually most effective when it is driven by the biological objective rather than by software familiarity alone.
In practical research settings, the first question is what kind of relationship the study is trying to resolve. If the goal is to identify specific cross-omics molecular pairs, such as transcript–protein concordance or enzyme–metabolite associations, feature-level correlation methods are usually the most direct starting point. If the goal is to characterize shared cross-omics structure distributed across many variables, latent-variable methods such as sparse CCA are generally more informative. When the study needs to distinguish shared biological signals from omics-specific variation, factor-model approaches such as MOFA+ provide a clearer framework. If the emphasis is on coordinated modules, hubs, or pathway-level organization, network methods such as WGCNA are often more appropriate. Finally, if the study is explicitly phenotype-driven and aims to derive a multi-omics biomarker panel or classifier, supervised methods such as DIABLO may be the better fit.
In most high-quality multi-omics studies, these method families are often combined to improve both robustness and interpretability. A common strategy is to begin with latent-variable or factor-based methods to identify stable shared structure across datasets, then apply network or feature-level analysis to refine biological interpretation and highlight specific molecules, modules, or pathways. In phenotype-oriented studies, supervised integration can be added after unsupervised structure discovery to prioritize features with both biological coherence and predictive relevance. This combined strategy is often more informative than relying on any single method in isolation because it allows different analytical layers to complement one another.
Table 1. Comparison of Major Method Families in Multi-Omics Correlation Analysis
| Method Family | Core Analytical Focus | Representative Method | Best Biological Question | Typical Output | Main Strengths | Main Limitations |
|---|---|---|---|---|---|---|
| Feature-level association | Pairwise co-variation between individual molecules across omics layers | Pearson/Spearman correlation | Which transcript–protein or protein–metabolite pairs change together? | Ranked molecular pairs, correlation matrices, filtered edge lists | Simple, intuitive, easy to visualize and interpret | High multiple-testing burden; indirect associations are common; limited ability to capture higher-order structure |
| Latent-variable integration | Shared variation axes across whole omics blocks | Sparse CCA | Is there a shared cross-omics structure linking two or more datasets? | Canonical components, selected cross-omics feature sets | Well suited to high-dimensional data; captures distributed multi-feature relationships | Sensitive to tuning, sample size, and preprocessing; component interpretation may require care |
| Factor-model decomposition | Shared versus omics-specific sources of variation | MOFA+ | Which signals are truly shared across omics layers, and which are layer-specific? | Shared factors, view-specific factors, factor loadings | Strong for disentangling common and private structure; useful in complex multimodal studies | Less intuitive than pairwise methods for readers unfamiliar with latent-factor models |
| Network or module analysis | Coordinated molecular groups, hubs, and cross-omics modules | WGCNA | Do molecules form co-regulated modules or pathway-level functional groups? | Modules, eigengenes, hub features, trait-associated subnetworks | Biologically interpretable; useful for pathway-scale discovery and visualization | Strong dependence on preprocessing, thresholding, and stability; visually rich networks can be misleading |
| Supervised multi-omics integration | Cross-omics feature selection guided by phenotype or class labels | DIABLO | Which multi-omics features best distinguish phenotypic groups or predict outcomes? | Multi-omics biomarker panels, discriminative components, classification signatures | Integrates omics structure with phenotype separation; useful for biomarker discovery | Requires predefined groups; more vulnerable to overfitting if validation is weak |
4. A Practical Workflow for Multi-Omics Correlation Analysis
A reliable multi-omics correlation analysis depends on more than choosing the right algorithm. In most projects, result quality is determined by sample matching, omics-specific preprocessing, feature selection, and validation just as much as by the integration model itself. A practical workflow therefore begins with study design, moves through data preparation and method selection, and ends with statistical and biological validation.
4.1 Define the Biological Objective and Integration Cohort
The first step is to clarify what the analysis is expected to deliver. Multi-omics correlation analysis may be used for candidate pair discovery, shared-structure analysis, biomarker selection, or module and hub identification. These goals require different method families and different validation strategies, so the primary objective should be fixed before modeling begins.
At the same time, the integration cohort must be defined clearly. Only samples that are truly comparable across omics layers should be included. In most correlation-driven studies, the safest starting point is a matched sample set with consistent phenotype labels, time points, and processing conditions.
Key output: a finalized sample annotation table and a clearly stated analysis goal.
4.2 Preprocess Each Omics Layer for Cross-Omics Integration
Transcriptomics, proteomics, and metabolomics should not be merged before they are individually cleaned and normalized. Each layer has its own data structure, noise pattern, and missing-value behavior, so preprocessing must be performed with omics-specific logic.
A practical minimum includes:
- removal of low-quality features and obvious outlier samples
- normalization and transformation appropriate to each platform
- review of batch effects and signal distribution
- cautious handling of missing values, especially in proteomics and metabolomics
Key output: one cleaned and analysis-ready matrix per omics layer.
4.3 Filter Features and Build the Integration Matrix
Integration becomes unstable when too many weak or low-information features are retained. Before running cross-omics models, it is usually helpful to filter features based on quality, variance, or biological relevance. For phenotype-driven studies, this may include preselecting informative candidates. For exploratory studies, broader feature sets may be retained, but low-confidence signals should still be removed.
Feature mapping should also be handled carefully. Gene–protein–metabolite relationships are rarely one-to-one, so pathway-level or module-level interpretation is often more robust than direct pair matching alone.
Key output: a documented feature set for integration, together with harmonized annotations.
4.4 Select the Integration Method and Generate Interpretable Outputs
Once the data are aligned and filtered, the primary method should be selected based on the research question. Feature-level correlation is useful for specific molecular pairs, sparse CCA for shared structure, MOFA+ for shared versus omics-specific variation, WGCNA for modules, and DIABLO for phenotype-guided feature selection.
The analysis should end with interpretable outputs rather than only model objects. Depending on the method, these may include ranked correlation pairs, shared components, latent factors, module summaries, hub features, or phenotype-linked multi-omics panels.
Key output: result tables and figures that can be directly interpreted biologically.
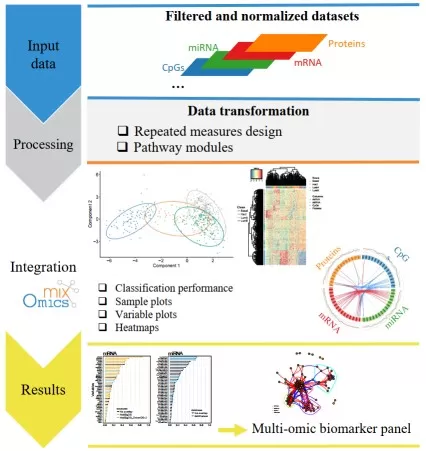
Figure 5. A standard DIABLO workflow. Image adapted from Singh et al. (2019), Bioinformatics, 35(17), 3055–3062.
4.5 Validate Results Statistically and Biologically
The final step is to test whether the results are stable and biologically meaningful. At minimum, this should include resampling, cross-validation, permutation testing, or sensitivity analysis where appropriate. Biological review is equally important: top signals should be checked against pathway context, known molecular relationships, and, when possible, independent datasets or targeted follow-up experiments.
In practice, the most credible result is rarely the longest list of associations. It is usually a smaller set of cross-omics signals that remains stable after validation and makes sense in biological context.
Key output: a refined set of robust, interpretable, and reportable findings.
5. Applications of Multi-Omics Correlation Analysis in Biomedical Research
Multi-omics correlation analysis is widely used in cancer research, metabolic disease, immunology, and other complex disease fields because it can connect molecular changes across transcriptomic, proteomic, and metabolomic layers within the same biological system. By revealing coordinated patterns that cannot be captured by a single omics dataset alone, it supports biomarker discovery, molecular subtype characterization, and pathway-level mechanism interpretation.
5.1 Cancer Biomarker Discovery and Pathway Stratification
Cancer is one of the clearest use cases for multi-omics correlation analysis because regulatory signals often become decoupled across DNA, RNA, protein, and phosphoprotein levels. A landmark breast cancer proteogenomics study showed that integrating genomic, transcriptomic, proteomic, and phosphoproteomic data can reveal pathway activity and regulatory consequences that are not obvious from mRNA-level analysis alone (Mertins et al., 2016). In this context, latent-variable methods and module-based approaches are particularly valuable because they can capture coordinated pathway behavior rather than relying only on one-gene-at-a-time differences.
Large consortium resources such as CPTAC have extended this logic by generating harmonized cross-omics datasets across multiple tumor types and making integrated interpretation more reproducible at scale (Li et al., 2023). For translational studies, this makes multi-omics correlation analysis especially useful for prioritizing biomarkers that remain coherent across molecular layers rather than appearing significant in only one assay.
5.2 Neurodegenerative Disease Mechanism and Molecular Stratification Studies
Neurodegenerative diseases are shaped by molecular heterogeneity across multiple regulatory layers, making them well suited to multi-omics correlation analysis. Integrated analysis can help connect cross-omics dysregulation with disease progression, pathological variation, and subtype-specific biology.
A representative example is a study on Alzheimer's disease that integrated transcriptomic, proteomic, metabolomic, and epigenomic data from brain and blood samples. The study established a unified multi-omics taxonomy, derived a molecular progression index, and resolved three robust disease subtypes linked to distinct pathological and clinical features (Iturria-Medina et al., 2022). This case shows how multi-omics integration can support molecular stratification and system-level interpretation in neurodegenerative disease research.
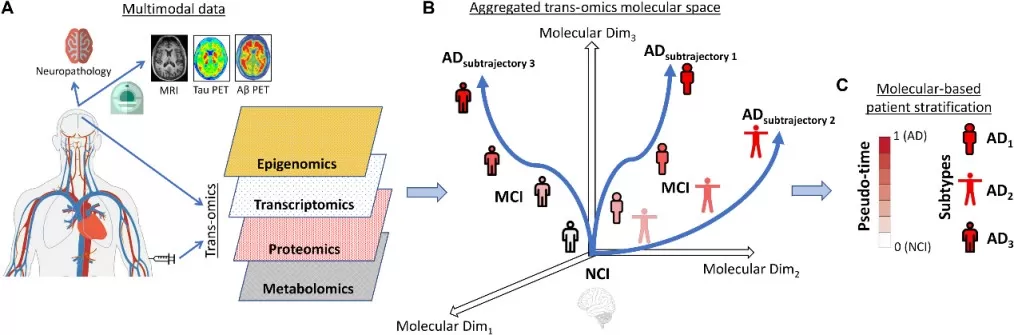
Figure 6. Schematic approach for multi-omics molecular integration and patient stratification in the late-onset AD dementia spectrum. Image adapted from Iturria-Medina et al. (2022), Science Advances, 8(46), eabo6764.
MetwareBio: The Right Partner for Your Multi-Omics Research
Reliable multi-omics correlation analysis depends on more than algorithm choice. It requires consistent sample handling, high-quality data generation across platforms, and bioinformatics support that extends from preprocessing and quality control to integration, interpretation, and visualization. MetwareBio supports this type of workflow through integrated capabilities in proteomics, metabolomics, lipidomics, and broader multi-omics analysis, together with the Metware Cloud Platform for downstream data exploration and visualization.
For multi-omics study design, sample requirements, or analysis support, contact MetwareBio for technical consultation.
Contact UsRead More: Multi-Omics Analysis Methods and Workflows
Multi-omics correlation analysis is one step in a broader integration pipeline. Explore these related articles to deepen your understanding of upstream statistical testing, pathway interpretation, and visualization strategies that complement cross-omics correlation.
Correlation heatmaps are the most common visual output of feature-level multi-omics analysis. Learn how to choose between Pearson and Spearman coefficients, design effective color scales, and interpret heatmap patterns to identify meaningful molecular relationships across omics datasets.
After identifying correlated cross-omics features, pathway enrichment analysis provides biological context. This guide compares over-representation analysis with gene set enrichment analysis to help you select the right approach for translating correlation results into pathway-level narratives.
Functional annotation databases provide the biological mapping needed to interpret cross-omics modules and latent factors. Understand when to use COG/KOG, GO, or KEGG for annotating correlated gene, protein, and metabolite sets in multi-omics integration studies.
Reactome offers a curated, pathway-centric alternative to KEGG for interpreting multi-omics results. Learn how Reactome's reaction-level detail can complement correlation-based module discovery and provide deeper mechanistic insight into coordinated omics signals.
References
- Liu, Y., Beyer, A., & Aebersold, R. (2016). On the Dependency of Cellular Protein Levels on mRNA Abundance. Cell, 165(3), 535–550. https://doi.org/10.1016/j.cell.2016.03.014
- Lancaster, S. M., Sanghi, A., Wu, S., & Snyder, M. P. (2020). A Customizable Analysis Flow in Integrative Multi-Omics. Biomolecules, 10(12), 1606. https://doi.org/10.3390/biom10121606
- Schober, P., Boer, C., & Schwarte, L. A. (2018). Correlation Coefficients: Appropriate Use and Interpretation. Anesthesia and Analgesia, 126(5), 1763–1768. https://doi.org/10.1213/ANE.0000000000002864
- de Winter, J. C., Gosling, S. D., & Potter, J. (2016). Comparing the Pearson and Spearman correlation coefficients across distributions and sample sizes: A tutorial using simulations and empirical data. Psychological Methods, 21(3), 273–290. https://doi.org/10.1037/met0000079
- Jiang, M. Z., Aguet, F., Ardlie, K., et al. (2023). Canonical correlation analysis for multi-omics: Application to cross-cohort analysis. PLoS Genetics, 19(5), e1010517. https://doi.org/10.1371/journal.pgen.1010517
- Bouhaddani, S. E., Houwing-Duistermaat, J., Salo, P., Perola, M., Jongbloed, G., & Uh, H. W. (2016). Evaluation of O2PLS in Omics data integration. BMC Bioinformatics, 17 Suppl 2(Suppl 2), 11. https://doi.org/10.1186/s12859-015-0854-z
- Singh, A., Shannon, C. P., Gautier, B., Rohart, F., Vacher, M., Tebbutt, S. J., & Lê Cao, K. A. (2019). DIABLO: an integrative approach for identifying key molecular drivers from multi-omics assays. Bioinformatics, 35(17), 3055–3062. https://doi.org/10.1093/bioinformatics/bty1054
- Witten, D. M., & Tibshirani, R. J. (2009). Extensions of sparse canonical correlation analysis with applications to genomic data. Statistical Applications in Genetics and Molecular Biology, 8(1), Article28. https://doi.org/10.2202/1544-6115.1470
- Argelaguet, R., Arnol, D., Bredikhin, D., Deloro, Y., Velten, B., Marioni, J. C., & Stegle, O. (2020). MOFA+: a statistical framework for comprehensive integration of multi-modal single-cell data. Genome Biology, 21(1), 111. https://doi.org/10.1186/s13059-020-02015-1
- Brown, B. C., Wang, C., Kasela, S., et al. (2023). Multiset correlation and factor analysis enables exploration of multi-omics data. Cell Genomics, 3(8), 100359. https://doi.org/10.1016/j.xgen.2023.100359
- Langfelder, P., & Horvath, S. (2008). WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics, 9, 559. https://doi.org/10.1186/1471-2105-9-559
- Mertins, P., Mani, D. R., Ruggles, K. V., et al. (2016). Proteogenomics connects somatic mutations to signalling in breast cancer. Nature, 534(7605), 55–62. https://doi.org/10.1038/nature18003
- Li, Y., Dou, Y., Da Veiga Leprevost, F., et al. (2023). Proteogenomic data and resources for pan-cancer analysis. Cancer Cell, 41(8), 1397–1406. https://doi.org/10.1016/j.ccell.2023.06.009
- Iturria-Medina, Y., Adewale, Q., Khan, A. F., et al. (2022). Unified epigenomic, transcriptomic, proteomic, and metabolomic taxonomy of Alzheimer's disease progression and heterogeneity. Science Advances, 8(46), eabo6764. https://doi.org/10.1126/sciadv.abo6764


