KEGG enrichment analysis is one of the most effective ways to turn omics data into biological insight. After identifying differentially expressed genes, proteins, or metabolites, the next question is no longer simply what changed, but what those changes mean. KEGG enrichment helps answer that question by showing which pathways are overrepresented in your dataset and therefore most likely to be involved in the biological response you are studying. Yet generating a KEGG enrichment table is only the first step. To draw meaningful conclusions, researchers need to understand how to interpret key metrics such as Gene Count, Rich Factor, and pathway significance—and how to read KEGG pathway maps in a biologically relevant way. In this article, we break down what these metrics really mean, how they complement one another, and how to move from a list of enriched pathways to a clearer, more mechanistic interpretation of your results.
1. WHAT KEGG ENRICHMENT ANALYSIS ACTUALLY TELLS YOU?
At its core, KEGG enrichment analysis asks a simple question: are the molecules in your input list—such as genes, proteins, or metabolites—found in a given pathway more often than expected by chance?
For example, imagine you have identified 300 differentially expressed genes from an RNA-seq experiment. KEGG enrichment analysis compares that list against annotated KEGG pathways and evaluates whether certain pathways contain more of those genes than would be expected if the list were random. If a pathway shows significant overrepresentation, it suggests that the biological process represented by that pathway may be involved in your experimental response.
However, enrichment analysis does not prove that the pathway is activated, inhibited, or causally driving the phenotype. It only tells you that the pathway is statistically overrepresented in your list. True biological interpretation depends on where the affected genes are located in the pathway, how strongly they change, whether they move in the same direction, and whether the result fits the broader experimental context.
That is why KEGG enrichment results should always be interpreted as evidence for pathway involvement, not as a direct functional conclusion.
2. HOW TO INTERPRET THE MAIN METRICS IN KEGG ENRICHMENT RESULTS?
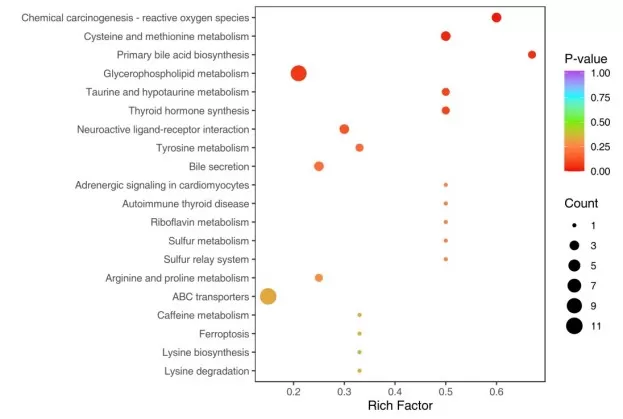
Figure 1. KEGG Enrichment Analysis Bubble Plot. Each bubble represents an enriched pathway; bubble size reflects Gene Count, x-axis position reflects Rich Factor, and color indicates statistical significance (p-value or FDR).
2.1 Gene Count
Gene Count is the number of genes from your input list that are annotated to a specific KEGG pathway. In other words, it represents the overlap between your dataset and that pathway. Although this metric is often labeled as "Gene Count", it may also represent proteins or metabolites, depending on the type of omics data and the annotation strategy used in the analysis.
This is usually the most intuitive metric. A pathway with 18 overlapping genes appears more substantial than one with 3. Gene Count is also useful as a practical reliability check: pathways supported by only one or two genes are often harder to interpret with confidence.
But Gene Count alone can be misleading. KEGG pathways vary greatly in size. Some pathways are broad and contain hundreds of genes, while others are small and tightly defined. A large pathway naturally has more opportunities to collect hits, even if only a small fraction of the pathway is actually affected. As a result, a high Gene Count does not necessarily mean the biological signal is focused or important.
So Gene Count answers one question well: how many of my genes fall into this pathway? What it does not answer is: how concentrated is the perturbation within that pathway?
2.2 Rich Factor
Rich Factor helps answer that second question. It is typically defined as:
Rich Factor = Gene Count / total number of genes annotated to that pathway
Unlike Gene Count, which is a raw number, Rich Factor is a proportion. It tells you how much of the pathway is represented by your input list. This makes Rich Factor especially useful for evaluating pathway specificity. A pathway with 10 changed genes out of 20 total genes has a Rich Factor of 0.50, meaning half of the pathway is affected. By contrast, a pathway with 20 changed genes out of 300 total genes has a Rich Factor of only 0.067. Although the second pathway has the larger Gene Count, the first one shows a much more concentrated disturbance. That distinction matters. Gene Count captures the absolute size of overlap, while Rich Factor captures the intensity or focus of enrichment.
In practice, pathways with both a reasonable Gene Count and a relatively high Rich Factor are often more biologically informative than pathways with a high count but very low proportional enrichment. Rich Factor is especially helpful when comparing pathways of very different sizes.
2.3 p-value
The p-value reflects the statistical significance of the overlap between your gene list and a pathway. It estimates how likely it would be to observe that overlap, or a more extreme one, if the genes were distributed randomly. In KEGG over-representation analysis, this is commonly calculated using Fisher's exact test or the hypergeometric test.
A small p-value suggests that the overlap is unlikely to be random. That makes it useful for identifying candidate pathways. However, the p-value has important limitations. First, it is influenced by pathway size, the size of your input list, and the background gene set used in the analysis. Second, it does not tell you whether the pathway is biologically central, mechanistically coherent, or directly relevant to your phenotype. A pathway can have an impressive p-value and still be too broad or generic to offer much interpretive value. For that reason, raw p-values should be treated as an initial statistical signal, not a standalone decision criterion.
2.4 Adjusted p-value / FDR
Because KEGG enrichment analysis tests many pathways at once, some pathways may appear significant purely by chance. This is known as the multiple testing problem. To account for it, enrichment results usually include an adjusted p-value, most commonly calculated using the Benjamini–Hochberg method. This adjustment controls the False Discovery Rate (FDR), which estimates the expected proportion of false positives among the pathways considered significant. As a result, the adjusted p-value—often reported as an FDR or q-value—is generally more reliable than the raw p-value for identifying meaningful pathways.
In practice, researchers typically prioritize pathways with FDR < 0.05, since these are more likely to represent robust signals rather than random findings. By contrast, pathways with low raw p-values but nonsignificant FDR values should be interpreted with caution.
Understanding p-value and Adjusted p-value (FDR) in KEGG Enrichment Analysis
| Metric | What It Means | Strength | Limitation | Practical Use |
|---|---|---|---|---|
| p-value | The probability that the observed pathway enrichment occurred by chance | Useful for identifying potentially enriched pathways | Does not correct for multiple testing | Use as an initial indicator of enrichment |
| Adjusted p-value / FDR | Shows whether a pathway is still significant after accounting for all the pathways tested | More reliable for assessing true enrichment | More stringent and may exclude weaker signals | Use as the main criterion for pathway significance |
2.5 Pathway Ranking
Most KEGG enrichment results are presented as ranked lists, but ranking is often misunderstood. The "top" pathway is only the top pathway according to the metric used by that specific tool or visualization. In one result table, pathways may be ranked by raw p-value. In another, they may be ranked by adjusted p-value. In bubble plots, the x-axis may represent Rich Factor while dot size represents Gene Count and color represents significance. This means pathway ranking is not universal. It depends on how the result was generated and displayed.
More importantly, the pathway ranked first is not necessarily the most biologically important one. It may simply be the largest pathway, the most statistically stable one, or the one that best fits the ranking rule used by the software. Ranking helps you navigate the results. It does not replace biological judgment.
3. WHY STATISTICALLY SIGNIFICANT PATHWAYS ARE NOT ALWAYS THE MOST IMPORTANT?
One of the most common mistakes in KEGG interpretation is assuming that the most significant pathway is automatically the most meaningful. In reality, statistical significance and biological importance are related but not identical.
A large pathway can become highly significant because it contains many genes and therefore collects overlap easily. Broad pathways such as "Metabolic pathways", "Pathways in cancer", "PI3K-Akt signaling pathway", or "MAPK signaling pathway" often rise to the top for this reason. These results are not necessarily wrong, but they may be too general to explain the specific biology in your system.
Another issue is that some pathways reflect common cellular responses rather than phenotype-specific mechanisms. Pathways related to ribosomes, oxidative phosphorylation, proteasome activity, or general stress responses may repeatedly appear in many datasets. They may indeed be affected, but they are not always the best starting point for mechanistic interpretation.
A particularly important warning sign is the combination of significant p-value but low Rich Factor. This often means the pathway is statistically enriched because it is large, while only a small proportion of its components are actually represented in your dataset. Such pathways may indicate background involvement rather than a focused biological shift.
In contrast, the most compelling pathways often show a combination of:
- a meaningful Gene Count
- a relatively high Rich Factor
- a robust FDR
- strong biological relevance to the experiment
The best interpretation usually comes from balancing all four.
4. A PRACTICAL FRAMEWORK FOR PRIORITIZING KEGG PATHWAYS
When reviewing KEGG enrichment results, a useful strategy is to read the table in the following order:
Step 1: Filter by adjusted significance. Start with pathways that pass a reasonable FDR threshold. This removes many weak or unstable signals.
Step 2: Check Gene Count. Exclude pathways supported by too few genes unless they are highly relevant biologically. Very small overlaps can be difficult to interpret confidently.
Step 3: Compare Rich Factor. Among statistically significant pathways, prioritize those with stronger proportional enrichment. This helps identify focused pathway perturbations rather than diffuse ones.
Step 4: Evaluate biological plausibility. Ask whether the pathway makes sense for your sample type, treatment, disease model, or phenotype. A pathway that fits the biology is usually more valuable than a statistically stronger but biologically generic one.
Step 5: Return to the pathway map. Once you have shortlisted candidate pathways, inspect their KEGG maps directly. This is where the interpretation becomes mechanistic rather than purely statistical.
5. WHY KEGG PATHWAY MAPS MATTER?
A KEGG pathway name is only a label. A KEGG pathway map is the actual biological context. Pathway maps show how genes, proteins, enzymes, compounds, and reactions are organized within a curated network. Instead of simply telling you that a pathway is enriched, the map shows where your altered genes sit in the pathway and how they relate to one another.
Before drawing biological conclusions from a KEGG pathway map, it is important to understand the basic visual language of the diagram. In these maps, nodes usually represent genes, proteins, or enzymes—most often shown as rectangles—while metabolites or other compounds are commonly depicted as circles. The edges, including arrows and connecting lines, indicate relationships such as activation, inhibition, phosphorylation, or metabolic flow. This visual framework allows researchers to see not only which components are altered, but also where those changes occur within the pathway and how they may influence downstream biological events. Because mapped omics figures often use customized highlighting schemes, the meaning of colors such as red, blue, or yellow should always be interpreted according to the figure legend rather than assumed to reflect a universal KEGG standard.
This makes a major difference. Two pathways may have identical Gene Counts and similar FDR values, yet one may show a tight cluster of changes around a key regulatory branch, while the other shows scattered hits across unrelated regions. Statistically, they may appear similar. Biologically, they are not.
When interpreting a KEGG pathway map, ask the following questions:
- Are the altered genes clustered in one branch or scattered across the map?
- Do they localize to upstream receptors, signaling intermediates, transcriptional outputs, or metabolic endpoints?
- Are they centered around a known regulatory hub or bottleneck enzyme?
- Do the changes suggest a coherent direction, such as pathway activation, repression, or rewiring?
- Are the affected nodes directly relevant to your phenotype?
These questions often reveal whether the enriched pathway represents a true mechanistic signal or only a loose statistical association.
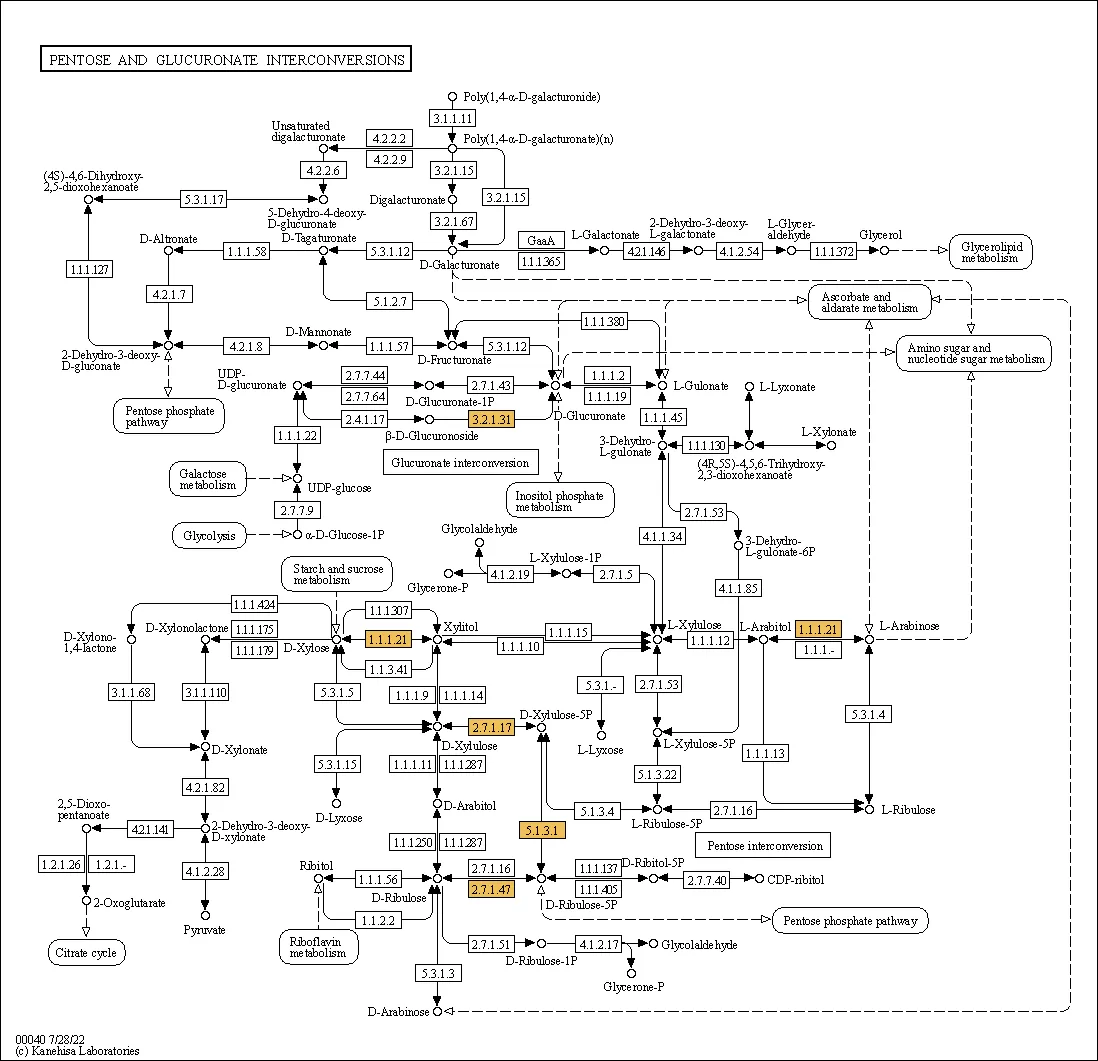
Figure 2. KEGG Pathway Map of Differentially Expressed Proteins. Highlighted nodes indicate proteins with significant expression changes; the map reveals their positions within the pathway network and potential downstream effects.
6. HOW PATHWAY MAPS IMPROVE INTERPRETATION: AN EXAMPLE
Consider a dataset in which "Pathways in cancer" appears among the top enriched terms. On its own, that result is too broad to be very informative. The pathway contains many signaling modules, including cell cycle, apoptosis, growth factor signaling, angiogenesis, and more. But once you map your genes onto the KEGG pathway diagram, a more specific picture may emerge. You might find that your altered genes cluster around the p53-MDM2-p21 axis, with little change in other parts of the cancer pathway. That pattern would suggest that the meaningful biology is not "cancer pathways" in a general sense, but rather p53-associated cell cycle arrest and apoptotic stress response.
This is exactly why pathway maps are essential. They convert a broad pathway label into a more precise mechanistic hypothesis.
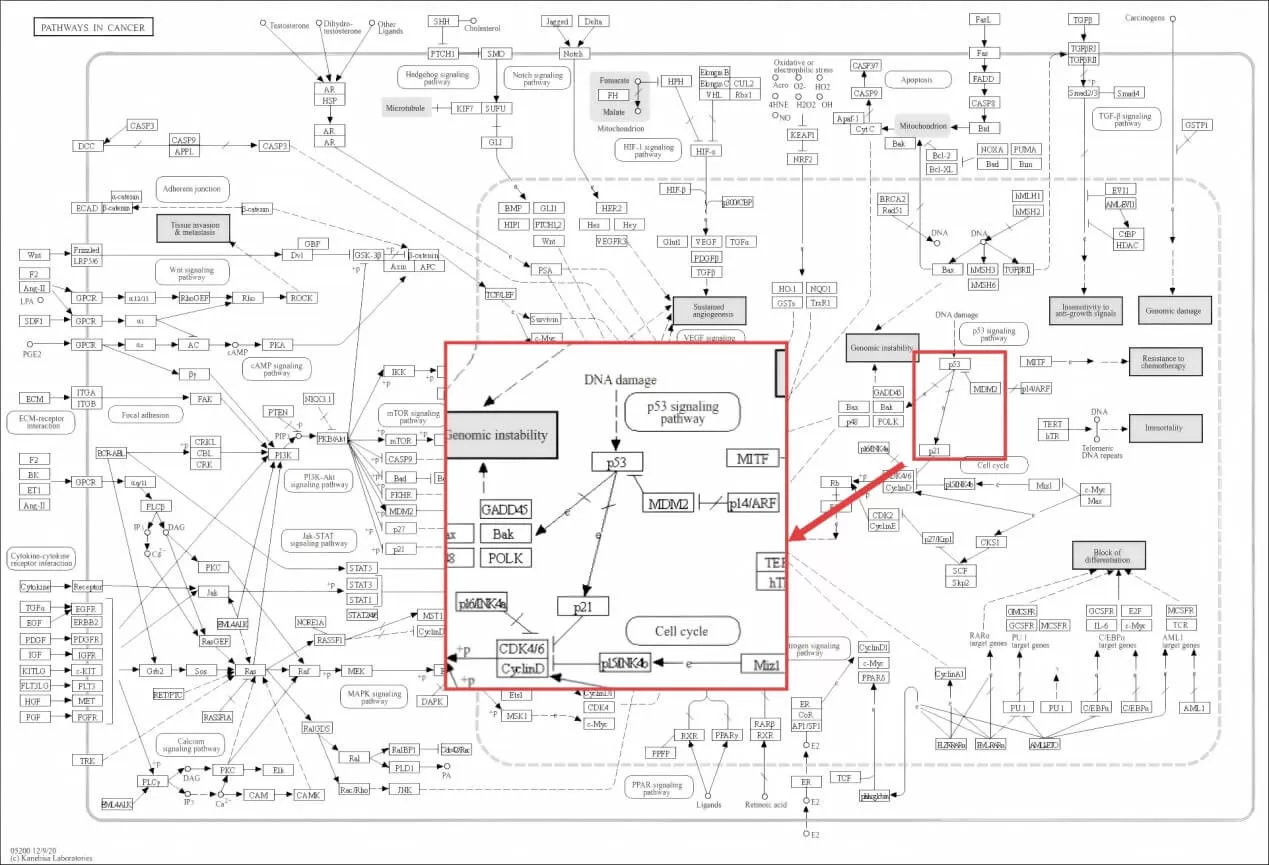
Figure 3. Zooming In on the p53–MDM2–p21 Axis in the KEGG "Pathways in Cancer" Map. By focusing on a specific regulatory module within a broad pathway, researchers can derive more precise mechanistic hypotheses from enrichment results.
7. HOW TO INTERPRET MULTIPLE ENRICHED PATHWAYS TOGETHER?
In most datasets, KEGG enrichment analysis does not highlight just one pathway, but a group of related pathways that overlap in function and collectively reflect a broader biological response. For that reason, pathways should not always be interpreted one by one in isolation. A more effective approach is to look for shared biological meaning across the enriched results and identify the larger process they point to.
For example, if your results include the T cell receptor signaling pathway, B cell receptor signaling pathway, NF-kappa B signaling pathway, cytokine-cytokine receptor interaction, and Th17 cell differentiation, it is often more informative to interpret them as parts of a broader biological theme rather than as separate findings. Together, these pathways suggest coordinated immune activation, particularly involving adaptive immune signaling, inflammatory regulation, and immune cell differentiation. In this case, the main insight is not simply that several pathways are enriched, but that they collectively support a more coherent biological interpretation.
This type of integrative reading is often more valuable than focusing too narrowly on individual pathway names. By moving from single enriched pathways to pathway clusters and then to a unifying biological theme, researchers can develop a clearer and more meaningful interpretation of their data.
| Common Pitfall | What It Means | Better Approach |
|---|---|---|
| Focusing only on the top-ranked pathway | Ranking is not the same as biological importance | Review significance, Rich Factor, Gene Count, and biological relevance together |
| Using raw p-values without considering FDR | Some pathways may look significant by chance | Prioritize adjusted p-values or FDR |
| Equating high Gene Count with high relevance | Large pathways tend to collect more hits | Interpret Gene Count together with pathway size and Rich Factor |
| Ignoring Rich Factor | Proportional enrichment may be overlooked | Use Rich Factor to assess how concentrated the signal is |
| Stopping at the enrichment table | Statistical results do not show pathway structure | Check KEGG pathway maps for gene positions and interactions |
| Interpreting pathways without biological context | Statistics alone cannot define relevance | Always relate enriched pathways to the phenotype and study design |
8. Conclusion: Turning KEGG Enrichment Analysis into Biological Insight
KEGG enrichment analysis is most valuable when it is interpreted beyond the ranked pathway list. Gene Count shows how many molecules map to a pathway, Rich Factor reflects how concentrated that overlap is, and adjusted p-value or FDR helps identify statistically reliable signals. But strong interpretation also depends on biological context, pathway structure, and whether related enriched pathways support the same mechanism. In the end, the goal of KEGG enrichment analysis is not simply to find significant pathways, but to build a clear, biologically meaningful explanation of your data.
MetwareBio: Multi-Omics Analysis Backed by Clear Biological Interpretation
MetwareBio provides proteomics, metabolomics, lipidomics, spatial metabolomics, and multi-omics analysis services to help researchers translate complex omics data into actionable insights. From KEGG enrichment analysis and pathway mapping to integrated biological interpretation, our team delivers reliable results with expert bioinformatics support.
Have a project in progress? Contact us to discuss your research goals.


