In MS-based proteomics, raw data should never be viewed as one-time experimental outputs. Instead, they are long-term digital assets that can continue to generate biological insight as reference databases improve, protein annotations expand, post-translational modifications (PTMs) are redefined, and computational workflows become more advanced. Under the search space, database completeness, and analytical standards available at the time of the original study, researchers often extract only the most accessible layer of information from proteomics raw files. When these same data are revisited using updated resources and improved analysis strategies, they may reveal peptides, proteins, and PTMs that were previously missed.
This is why proteomics raw data reanalysis is becoming an increasingly important strategy in modern mass spectrometry research. Datasets that once appeared fully interpreted may support entirely new biological conclusions when examined in a new computational context. This article focuses on the core concept of re-searching proteomics raw data and addresses three practical questions: what re-searching actually means in MS-based proteomics, when old raw data should be revisited, and what kinds of new findings such reanalysis can reveal.
1. What Does Re-Searching MS-Based Proteomics Raw Data Mean?
In MS-based proteomics, re-searching does not simply mean revisiting old identification tables or reinterpreting previously reported proteins. Rather, it means returning to the original raw files and performing peptide-spectrum matching, protein inference, statistical control, and downstream interpretation again under updated analytical conditions. These updated conditions may include improved protein sequence databases, expanded PTM settings, revised search parameters, or newer identification algorithms.
In practice, this process typically involves re-matching MS/MS spectra to theoretical peptide sequences, reapplying false discovery rate (FDR) control and filtering criteria, and generating a new set of peptide, protein, and sometimes quantitative results. For this reason, re-searching is best understood as a full proteomics data reanalysis workflow rather than a secondary interpretation of prior outputs. Published studies have already shown that large public proteomics datasets can be re-identified and re-quantified directly from raw files, producing results that differ from and may be more comprehensive than the original analysis. These studies provide clear methodological and practical support for the value of proteomics raw data reanalysis (Dai et al., 2024).
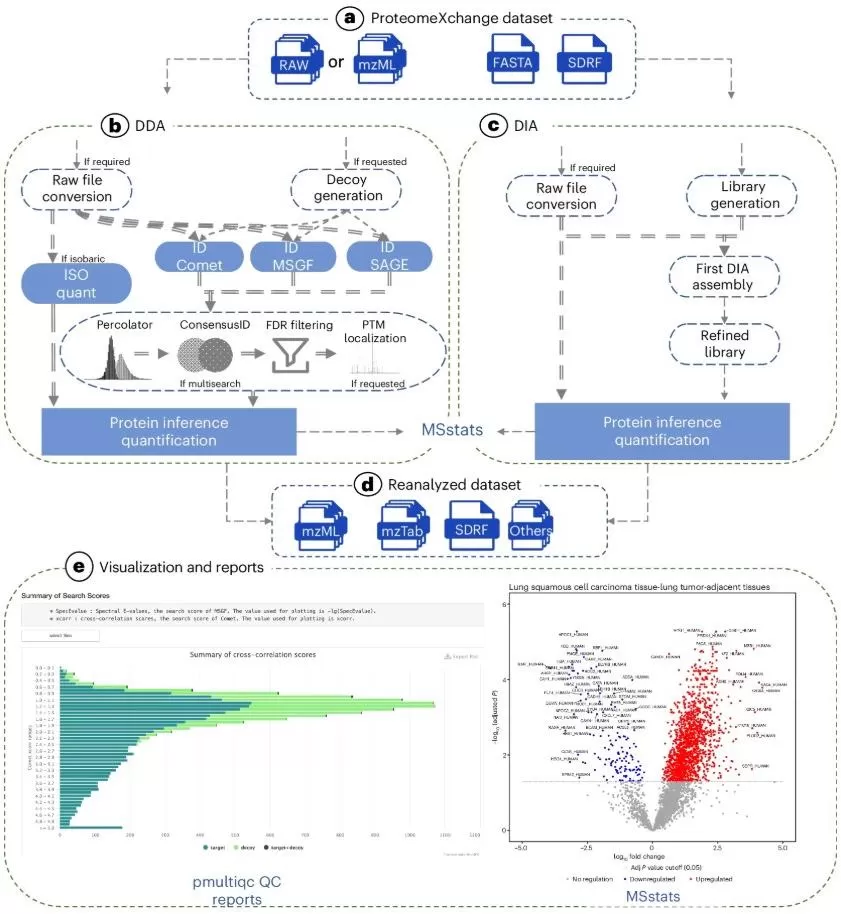
Figure 1. Workflow of a quantitative proteomics analysis platform capable of re-analyzing public proteomics data. Image reproduced from Dai et al., 2024, Nature Methods, 21(9), 1603–1607.
2. When Should You Re-Search MS-Based Proteomics Raw Data?
In proteomics, reanalysis is most valuable when the biological question remains relevant but the analytical context has changed. Even when the raw spectra are unchanged, improvements in databases, PTM knowledge, quality control strategies, and search engines can substantially alter peptide and protein identification outcomes. For researchers working with legacy LC-MS/MS datasets, the key question is not whether the data are old, but whether the current analytical framework is more informative than the original one.
2.1 New Proteomics Databases and Better Reference Sequences
Database updates are one of the most common and most defensible reasons to re-search old proteomics raw data. This is especially true in MS-based proteomics projects involving non-model organisms, incomplete reference proteomes, or rapidly evolving annotation resources. In many early studies, unidentified spectra were not necessarily uninformative; rather, the relevant peptide sequences may simply have been absent from the database used at the time.
This situation becomes even more important when researchers begin to use more specific search resources, such as strain-level databases, pangenome databases, MAG-derived protein sets, or updated UniProt and RefSeq releases. In these settings, proteomics raw data reanalysis can increase the number of confidently identified peptides, improve protein sequence coverage, and sometimes reveal previously unrecognized proteins or biologically relevant sequence variants. Several public proteomics reanalysis studies have shown that introducing newer or more targeted databases can substantially expand identification depth without any additional experimental cost (Levitsky et al., 2023); (Stroggilos et al., 2024). In other words, better sequence knowledge often unlocks new biological value from the same raw MS data.
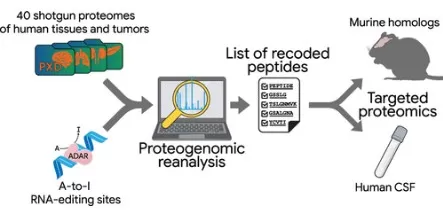
Figure 2. Re-analysis of 40 public shotgun proteomics datasets from different human tissues, focusing on the search for ADAR-mediated RNA editing events. Image reproduced from Levitsky et al., 2023, Journal of Proteome Research, 22(6), 1695–1711.
2.2 New PTMs and Updated Modification Priorities
In proteomics database searching, identification results depend strongly on which modifications are included in the search space. If the original study considered only a limited set of common variable modifications, then a substantial portion of MS/MS spectra may have remained uninterpreted simply because the relevant PTMs were not part of the analytical design. As the field evolves, researchers often shift from focusing only on standard modifications such as oxidation or acetylation to investigating newer or biologically emerging PTMs, including lactylation, crotonylation, glycosylation, or chemically induced modifications.
Reanalysis under updated PTM assumptions can therefore transform the interpretation of legacy proteomics datasets. Public reanalysis studies of global proteomics and phosphoproteomics data have shown that introducing targeted modification settings during re-searching can recover large numbers of modified peptides that were not reported in the original workflow (Hu et al., 2018). These findings highlight a key principle in MS-based proteomics: many previously unused spectra are not random noise, but signals that were missed because the original search parameters did not reflect the full biological or chemical complexity of the sample.
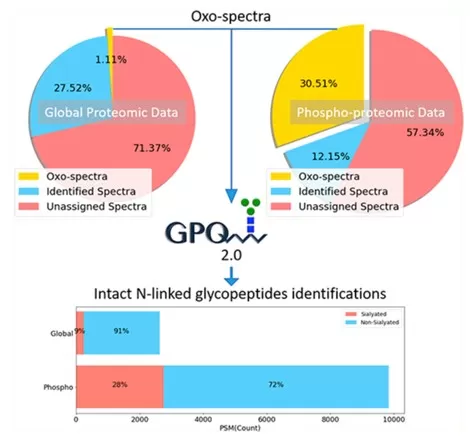
Figure 3. Re-analysis of conventional proteomics and phosphoproteomics data identified a large number of glycopeptides. Image reproduced from Hu et al., 2018, Analytical Chemistry, 90(13), 8065–8071.
2.3 Improved Search Parameters and QC Criteria
Another major reason to revisit legacy proteomics raw data is that search parameters and quality control standards often improve over time. Earlier MS-based proteomics studies sometimes emphasized identification yield, which could encourage the use of wider precursor or fragment mass tolerances, higher FDR thresholds, or relatively permissive filtering strategies. Although such settings may increase the total number of reported identifications, they can also elevate the risk of false positives and reduce the robustness of downstream biological interpretation.
Re-searching under more appropriate parameter settings allows researchers to reassess identification confidence using current best practices. In proteomics, this can be particularly important when the downstream goals include pathway analysis, protein network interpretation, biomarker prioritization, or quantitative modeling. Cleaner peptide and protein tables often support stronger biological conclusions, even if the total number of identifications becomes slightly smaller. From an analytical perspective, reanalysis in this context is not about maximizing output counts, but about improving data quality, reproducibility, and interpretability across the full proteomics workflow.
2.4 Advances in Search Algorithms and Identification Strategies
Proteomics raw data also become more valuable when the computational methods used to interpret them improve. Over the past several years, MS-based proteomics has seen major advances in peptide identification, including machine learning-enhanced scoring, open and semi-open search strategies, spectral library searching, and predicted spectrum-assisted matching. These developments have meaningfully changed how LC-MS/MS data are processed and how borderline or low-abundance spectra are evaluated.
As a result, reanalysis with modern search engines can recover peptides and proteins that were difficult to detect using earlier workflows, while also correcting some misidentifications from legacy analyses. In large public reanalysis efforts, different search strategies often produce structurally different peptide and protein identification results, even when applied to the same raw files. This observation reinforces an important point for proteomics researchers: the biological value of raw data is partly determined by the algorithmic tools available at the time of analysis. When those tools improve, revisiting old proteomics datasets can become scientifically justified and analytically rewarding (Dai et al., 2024); (Yang et al., 2023).
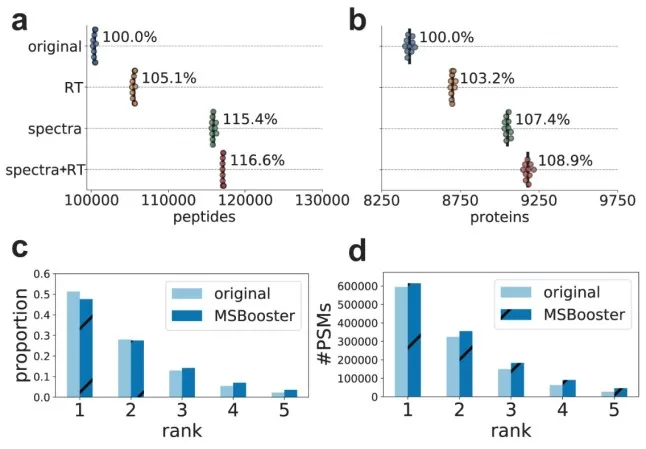
Figure 4. Deep learning-derived features from the new algorithm MSBooster improved peptide and protein identification rates by 16.6% and 8.9%, respectively. Image reproduced from Yang et al., 2023, Nature Communications, 14(1), 4539.
3. What Can MS-Based Proteomics Raw Data Reanalysis Reveal?
In proteomics, the value of reanalysis is ultimately measured by what new biological or analytical insight it can generate. Re-searching legacy raw data does not only improve peptide-spectrum matching statistics; it can also reshape protein-level interpretation, refine published conclusions, and create new opportunities for hypothesis-driven research. For this reason, proteomics raw data reanalysis should be viewed not merely as a technical exercise, but as a practical strategy for expanding the scientific return of existing MS datasets.
3.1 New Proteins, Peptides, and PTMs
The most immediate benefit of proteomics raw data reanalysis is the recovery of new molecular identifications. When legacy LC-MS/MS datasets are searched against updated reference proteomes, broader PTM settings, or more advanced search algorithms, spectra that were previously unmatched may now be assigned to specific peptides. At the protein level, this can improve sequence coverage, support the detection of low-abundance proteins, and reveal protein variants or newly annotated gene products that were absent from earlier analyses. At the modification level, reanalysis may also identify PTM-containing peptides that were originally missed because the relevant modification was not included in the search space.
These gains are especially important in MS-based proteomics because biological interpretation depends heavily on identification depth and confidence. A newly identified peptide can strengthen evidence for a protein, refine functional annotation, or point to an unexpected regulatory mechanism. Likewise, newly recovered PTMs may open the door to more precise mechanistic interpretation. In this sense, proteomics reanalysis does not just add more IDs; it expands the molecular resolution at which a biological system can be understood.
3.2 Refining and Validating Existing Conclusions
Not all proteomics raw data reanalysis is aimed at discovering something completely new. In many cases, its greatest value lies in testing how robust previously reported conclusions remain under improved analytical conditions. When older datasets are reprocessed using stricter FDR control, more appropriate filtering, or updated database resources, researchers can evaluate whether the original biological interpretation still holds. This is particularly important in MS-based proteomics studies that support mechanistic claims, pathway enrichment results, biomarker selection, or differential protein expression patterns.
Reanalysis can therefore serve as a form of analytical validation. Some conclusions may become stronger because the same biological signal remains detectable despite more rigorous processing. Other conclusions may prove highly sensitive to database choice, PTM settings, or search engine behavior, suggesting that they should be interpreted more cautiously. This kind of reassessment improves the reliability, reproducibility, and transparency of proteomics research. Rather than undermining earlier work, it often helps distinguish durable biological findings from results that depended too heavily on the limitations of the original computational workflow.
3.3 Reusing Raw Data for New Scientific Questions
Another major advantage of legacy proteomics data is that the same raw files can be repurposed to address new scientific questions that were not part of the original study design. As research priorities shift, previously collected MS-based proteomics datasets may become highly valuable for pilot analyses, exploratory validation, or cross-project integration. For example, a dataset initially generated for global protein profiling might later be reanalyzed for PTM discovery, alternative protein sequence evidence, proteogenomic interpretation, or benchmarking of a newly developed search workflow.
This flexibility makes proteomics raw files a reusable scientific resource rather than a fixed historical record. Because sample collection, protein extraction, LC-MS/MS acquisition, and large-scale biological experiments are often expensive and time-consuming, the ability to revisit existing raw data can substantially improve research efficiency. Reanalysis is also well suited to public repository reuse, where deposited proteomics datasets can support questions far beyond the scope of the original publication. In this way, MS-based proteomics raw data can continue to generate value across time, projects, and analytical frameworks, provided that researchers are willing to ask new questions of old spectra.
Turn Legacy Proteomics Data into New Insights
Old proteomics raw files may still contain biological signals that were missed the first time around. If you are revisiting archived LC-MS/MS datasets, MetwareBio offers proteomics data reanalysis support to help recover new peptides, proteins, and PTMs, refine existing conclusions, and unlock more value from data you already have. Our quantitative proteomics services include advanced bioinformatics pipelines and expert support for re-searching legacy datasets with updated databases, expanded PTM settings, and modern search algorithms.
Contact us to discuss your project and find the right reanalysis strategy.
Contact UsReferences
- Dai, C., Pfeuffer, J., Wang, H., Zheng, P., Käll, L., Sachsenberg, T., Demichev, V., Bai, M., Kohlbacher, O., & Perez-Riverol, Y. (2024). quantms: a cloud-based pipeline for quantitative proteomics enables the reanalysis of public proteomics data. Nature Methods, 21(9), 1603–1607. https://doi.org/10.1038/s41592-024-02343-1
- Levitsky, L. I., Ivanov, M. V., Goncharov, A. O., Kliuchnikova, A. A., Bubis, J. A., Lobas, A. A., Solovyeva, E. M., Pyatnitskiy, M. A., Ovchinnikov, R. K., Kukharsky, M. S., Farafonova, T. E., Novikova, S. E., Zgoda, V. G., Tarasova, I. A., Gorshkov, M. V., & Moshkovskii, S. A. (2023). Massive Proteogenomic Reanalysis of Publicly Available Proteomic Datasets of Human Tissues in Search for Protein Recoding via Adenosine-to-Inosine RNA Editing. Journal of Proteome Research, 22(6), 1695–1711. https://doi.org/10.1021/acs.jproteome.2c00740
- Stroggilos, R., Tserga, A., Zoidakis, J., Vlahou, A., & Makridakis, M. (2024). Tissue proteomics repositories for data reanalysis. Mass Spectrometry Reviews, 43(6), 1270–1284. https://doi.org/10.1002/mas.21860
- Hu, Y., Shah, P., Clark, D. J., Ao, M., & Zhang, H. (2018). Reanalysis of Global Proteomic and Phosphoproteomic Data Identified a Large Number of Glycopeptides. Analytical Chemistry, 90(13), 8065–8071. https://doi.org/10.1021/acs.analchem.8b01137
- Yang, K. L., Yu, F., Teo, G. C., Li, K., Demichev, V., Ralser, M., & Nesvizhskii, A. I. (2023). MSBooster: improving peptide identification rates using deep learning-based features. Nature Communications, 14(1), 4539. https://doi.org/10.1038/s41467-023-40129-9


