Matrix-Assisted Laser Desorption/Ionization Spatial Metabolomics (MALDI-MSI) has emerged as a transformative technology in metabolomics research, enabling direct visualization of metabolites, lipids, and small molecules within tissue sections. A critical factor that dictates the success of these experiments is the choice of matrix. The matrix not only facilitates analyte ionization but also impacts signal intensity, signal-to-noise ratio, spatial resolution, and overall data reproducibility. This article offers a systematic guide for choosing the most suitable MALDI matrix, highlighting the strengths, limitations, and practical applications of four key matrices — DHB, CHCA, 9-AA, and DAN. It also provides a stepwise decision framework to help researchers design robust spatial metabolomics experiments and maximize the interpretability of their data.
1. THE ROLE OF THE MATRIX IN MALDI IMAGING
In MALDI spatial metabolomics, the matrix is not merely a supporting reagent added to enable analysis. It is one of the most important determinants of ionization performance, molecular coverage, image quality, and data reliability throughout the entire MALDI-MSI workflow. In practical terms, the matrix acts as a molecular energy mediator between the laser and the analytes embedded in the tissue section. Without a suitable matrix, many metabolites and lipids cannot be efficiently desorbed and ionized, and high-quality spatial metabolomics data would be difficult to obtain.
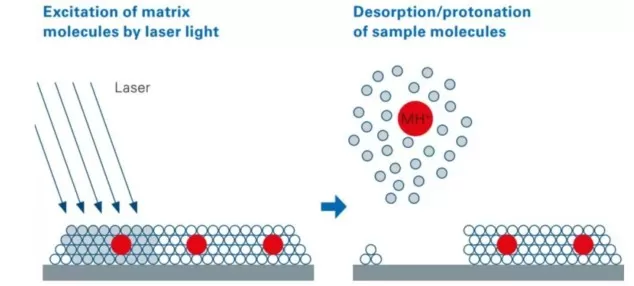
Figure 1. Mechanism of Matrix-Assisted Laser Desorption/Ionization (MALDI). The matrix absorbs UV laser energy, facilitating desorption and ionization of co-crystallized analyte molecules from the tissue surface.
A MALDI matrix is typically a low-molecular-weight organic compound that is uniformly deposited onto the surface of a tissue section before mass spectrometry imaging. During laser irradiation, the matrix strongly absorbs ultraviolet energy at the MALDI laser wavelength, undergoes rapid excitation and desorption, and assists in carrying nearby analyte molecules into the gas phase. At the same time, it facilitates proton transfer, cation transfer, or deprotonation, depending on the ion mode and the chemical properties of the analytes. Through this process, metabolites, lipids, and other small molecules are converted into gas-phase ions that can then be detected by the mass spectrometer.
An effective matrix must meet multiple technical criteria. It should:
- Exhibit strong UV absorption in the laser wavelength range used for MALDI, so that it can efficiently capture laser energy and initiate the desorption/ionization process. Insufficient UV absorption usually leads to weak ion generation and reduced detection sensitivity.
- Promote co-crystallization with target analytes for efficient desorption, because uniform and intimate mixing between the matrix and metabolites is essential for reproducible ionization across the tissue surface and for maintaining consistent signal quality.
- Maintain chemical stability under high vacuum conditions, ensuring that the matrix remains suitable throughout data acquisition and does not introduce instability, signal drift, or unwanted artifacts during MALDI-MSI analysis.
- Minimize background noise to avoid interference with low-abundance metabolites, especially in the low-mass range, where matrix-derived ions can easily suppress or obscure biologically relevant small-molecule signals.
2. COMMON MATRICES FOR MALDI SPATIAL METABOLOMICS
Over decades of MALDI-MS development, hundreds of matrix compounds have been explored. In practice, however, only a select few have become standard for spatial metabolomics due to their reproducibility, ionization efficiency, and compatibility with tissue imaging. Below is a detailed overview of the most widely adopted matrices.
2.1 2,5-Dihydroxybenzoic Acid (DHB)
DHB is arguably the most versatile matrix in the MALDI-MSI toolkit. It forms relatively homogeneous crystals and excels at detecting lipids, carbohydrates, and small metabolites in positive ion mode. Its hydrophilic nature and "cool" matrix characteristics — meaning it imparts less internal energy to analytes — reduce unwanted fragmentation. This makes DHB the default choice for lipid imaging and broad-spectrum metabolic screening.
- Ionization mode: Primarily positive ion mode.
- Target analytes: Lipids (including phosphatidylcholine [PC], sphingomyelin [SM]), carbohydrates, and small metabolites.
- Advantages: Offers high sensitivity for choline-containing lipids; robust crystallization that supports reproducible imaging.
- Applications: The matrix of choice for lipidomics studies in positive ion mode, particularly for profiling phospholipids across tissues.
- Limitations: DHB may generate heterogeneous crystal layers, limiting ultra-high spatial resolution; careful deposition or recrystallization may be needed for fine-scale imaging.
2.2 α-Cyano-4-hydroxycinnamic Acid (CHCA)
CHCA stands as the matrix of choice for peptides, amino acids, and small organic acids. It forms finer crystals than DHB (typically 10–15 μm), enabling higher spatial resolution imaging. CHCA is considered a "hotter" matrix — it transfers more internal energy to analytes — which can enhance ionization efficiency for certain compounds but may increase fragmentation risk for fragile metabolites. A 2023 study on brain tumor metabolomics found that recrystallized CHCA with positive polarity offered superior performance for both metabolite coverage and consistency with orthogonal techniques.
- Ionization mode: Positive ion mode.
- Target analytes: Amino acids, small metabolites, and peptides.
- Advantages: Supports ultrafine spatial resolution, enabling cellular-level metabolite mapping. Studies have shown that recrystallized CHCA provides superior sensitivity for metabolites in brain tumor tissues and aligns well with LC-MS and RNA-seq datasets.
- Applications: Cell heterogeneity studies, tumor microenvironment mapping, and any research where high spatial resolution is critical.
- Limitations: Less effective for highly acidic metabolites; deposition uniformity is essential to avoid signal variability.
2.3 9-Aminoacridine (9-AA)
9-AA excels in negative ion mode, offering exceptional sensitivity for acidic compounds.
- Ionization mode: Negative ion mode.
- Target analytes: Organic acids, nucleotides, phosphorylated metabolites, ATP/ADP, and energy-related metabolites.
- Advantages: High ionization efficiency for fatty acids, bile acids, and phosphorylated small molecules.
- Applications: Energy metabolism profiling, organic acid analysis, and nucleotide quantification.
- Limitations: Limited positive ion response; best used when acidic metabolite coverage is a priority.
2.4 1,5-Diaminonaphthalene (DAN)
DAN is an emerging matrix capable of dual-mode ionization, providing a "one-matrix, two-modes" solution.
- Ionization mode: Both positive and negative ion modes.
- Target analytes: Broad coverage of lipid classes, metabolites, and small molecules.
- Advantages: Enables sequential acquisition of positive and negative ion data from a single tissue section, enhancing information density. Sublimation techniques allow for uniform coatings below 20 μm, supporting high-resolution imaging.
- Applications: Ideal for rare or limited samples, comprehensive lipidomics studies, and experiments where maximizing molecular coverage is critical.
- Limitations: Requires careful control during deposition to ensure reproducibility.
2.5 DHB vs CHCA vs 9-AA vs DAN: Comparative Guide
To provide a clear and practical reference, the following table summarizes the key characteristics of the four most widely used MALDI matrices: DHB, CHCA, 9-AA, and DAN. It highlights their preferred ionization modes, target analyte classes, spatial resolution capabilities, and main advantages.
| Matrix | Ion Mode | Target Analytes | Spatial Resolution | Key Advantages |
|---|---|---|---|---|
| DHB | Positive | Lipids, carbohydrates, small metabolites | Moderate (>20 μm) | Broad applicability, strong response for choline-containing lipids |
| CHCA | Positive | Amino acids, small metabolites | High (<10 μm) | Fine crystals, supports high-resolution imaging |
| 9-AA | Negative | Organic acids, nucleotides, phosphorylated metabolites | Moderate (10–50 μm) | Excellent sensitivity for acidic metabolites |
| DAN | Positive & Negative | Lipids, small metabolites | High (<20 μm with sublimation) | Dual-mode capability, broad molecular coverage from single section |
3. STEPWISE FRAMEWORK FOR MATRIX SELECTION IN MALDI-MSI ANALYSIS
Matrix selection should be guided by three key considerations: the chemical nature of target analytes, desired spatial resolution, and sample availability. The chemical properties of metabolites, lipids, and small molecules determine which matrices can efficiently ionize them in either positive or negative ion mode. Spatial resolution requirements influence matrix choice because crystal size and deposition uniformity affect how precisely molecules can be mapped within the tissue. Finally, sample availability is critical when working with limited or precious tissue; a matrix that allows dual-mode acquisition or maximal molecular coverage can provide more information from a single section, increasing data yield while minimizing sample consumption. Taking these three factors into account provides a clear framework for selecting the most suitable matrix, which can then be systematically applied using a stepwise decision strategy.
3.1 Step 1: Identify Target Analytes
The first step in matrix selection is to clearly define the primary metabolites, lipids, or small molecules of interest. Different analyte classes ionize with varying efficiencies depending on the matrix chemistry and ionization mode. For instance, phospholipids generally respond better in positive ion mode with DHB, while acidic metabolites such as organic acids or nucleotides are more efficiently detected using negative ion matrices like 9-AA. Aligning the matrix choice with analyte chemistry ensures optimal ionization, enhanced signal intensity, and reliable metabolite detection across the tissue section. The following table provides a summary of recommended matrices, ion modes, and target analytes based on the chemical properties outlined above.
| Target Analytes | Recommended Matrix | Ion Mode |
|---|---|---|
| Phospholipids (PC, SM) | DHB | Positive |
| Phosphatidylethanolamine (PE), Phosphatidylinositol (PI) | DAN | Negative |
| Acidic small molecules (organic acids, nucleotides) | 9-AA | Negative |
| Amino acids, small metabolites | CHCA | Positive |
3.2 Step 2: Determine Spatial Resolution Needs
Spatial resolution considerations are critical for high-quality MALDI-MSI. Large or heterogeneous matrix crystals can reduce imaging precision, whereas fine, uniform deposition supports subcellular-level mapping. For resolutions above 50 μm, most matrices are suitable, but for 10–50 μm or sub-10 μm imaging, matrices with fine crystallization properties such as CHCA or sublimated DAN are preferred. Selecting the matrix in accordance with spatial resolution needs preserves the accurate spatial distribution of metabolites and enhances interpretability in tissue metabolomics studies. In summary:
- >50 μm: Most matrices are suitable; analyte compatibility is the primary factor.
- 10–50 μm: Use matrices with fine crystals such as CHCA or sublimated DAN/9-AA.
- <10 μm: CHCA is recommended for ultrafine imaging; its crystal structure supports subcellular resolution.
3.3 Step 3: Assess Sample Scarcity
Sample scarcity and tissue availability often influence matrix choice. For limited or precious samples, a matrix that allows dual-mode acquisition, such as DAN, can maximize molecular coverage by collecting both positive and negative ion data from the same section. This approach enhances information yield while minimizing sample consumption. Considering sample availability ensures that experimental design is efficient and that every section produces biologically meaningful, reproducible data suitable for comprehensive spatial metabolomics analysis.
4. ADVANCED CONSIDERATIONS FOR HIGH-QUALITY MALDI SPATIAL METABOLOMICS
Beyond basic matrix selection, several experimental factors play a critical role in determining the quality, reproducibility, and interpretability of MALDI spatial metabolomics data:
- Matrix Deposition Method: Techniques such as sublimation or automated matrix spraying can improve coating uniformity, reduce crystal size heterogeneity, and minimize analyte delocalization, thereby enhancing spatial fidelity and detection sensitivity.
- Co-Crystallization Optimization: Adjusting solvent composition, recrystallization conditions, and deposition parameters ensures that metabolites are evenly incorporated into the matrix layer. This step is particularly vital for small molecules with low ionization efficiency, as it directly impacts signal intensity and reproducibility across pixels.
- Instrument Parameters: Laser energy, spot size, raster step size, and acquisition speed should be matched to matrix properties and tissue type to achieve the desired balance between spatial resolution and signal intensity. Improper settings can lead to uneven ionization or signal suppression, compromising both qualitative and quantitative analyses.
- Data Integration: Data integration strategies enhance the biological interpretability of spatial metabolomics results. Combining MALDI-MSI with orthogonal approaches, such as LC-MS metabolomics or transcriptomics, can confirm metabolite identities and validate spatial patterns.
By considering these advanced factors in combination with matrix selection, researchers can generate robust, high-resolution, and biologically meaningful imaging datasets suitable for detailed tissue-level metabolite mapping.
5. CONCLUSION AND FUTURE PERSPECTIVES IN MALDI MATRIX DEVELOPMENT
In MALDI spatial metabolomics, there is no universal "best matrix." DHB, CHCA, 9-AA, and DAN each offer unique advantages and trade-offs. The ideal choice depends on the specific research question, target metabolite classes, spatial resolution requirements, and sample availability. By following a systematic matrix selection framework and optimizing deposition strategies, researchers can generate high-quality, reproducible imaging data that unlocks new biological insights.
Looking forward, advances in matrix chemistry and deposition technologies are poised to expand the capabilities of MALDI imaging. Novel sublimable matrices and hybrid organic–inorganic compounds offer improved sensitivity, reduced background noise, and dual-mode ionization for broader metabolite coverage. Automated and high-precision deposition systems are enabling ultra-fine crystal formation, supporting subcellular resolution imaging. As these innovations continue, MALDI spatial metabolomics will increasingly allow comprehensive, high-resolution, and quantitative tissue metabolite mapping, facilitating new insights in tumor biology, neuroscience, and systems-level metabolomics research.
MetwareBio: Your Trusted Partner for MALDI Spatial Metabolomics Analysis
MetwareBio offers advanced spatial metabolomics services powered by state-of-the-art MALDI imaging mass spectrometry technology. Through optimized matrix selection and deposition strategies, we deliver high-resolution, reproducible spatial mapping of metabolites, lipids, and other small molecules across diverse tissue samples.
From cancer biology to neuroscience and drug distribution studies, our service helps researchers uncover spatial molecular insights with confidence.
Need spatial metabolomics analysis or MALDI-MSI matrix optimization support? Contact MetwareBio to start your project.
Contact UsReferences
- Lu T, et al. Matrix Selection for the Visualization of Small Molecules and Lipids in Brain Tumors Using Untargeted MALDI-TOF Mass Spectrometry Imaging. Metabolites. 2023;13(11):1139.
- Thomas A, et al. Sublimation of new matrix candidates for high spatial resolution imaging mass spectrometry of lipids: enhanced information in both positive and negative polarities after 1,5-diaminonapthalene deposition. Anal Chem. 2012;84(4):2048–2054.


