Differential feature screening in omics research is more complex than simply ranking candidates by p-value. In a single untargeted experiment, thousands of metabolites or proteins are tested simultaneously, and the choice of metrics — Fold Change, p-value, FDR, or VIP — can dramatically alter which features make the final shortlist. This practical guide explains what each metric actually measures, when to use them in combination, and why the most defensible candidates are rarely those with the smallest nominal p-value. Through examples drawn from metabolomics, proteomics biomarker screening, and spatial omics differential analysis, we show how a layered, evidence-based screening framework produces results that are biologically meaningful, statistically credible, and worth the cost of downstream validation.
1. WHEN THREE ANALYSTS PRODUCE THREE DIFFERENT CANDIDATE LISTS
Suppose analyst A applies VIP>1 and identifies 60 metabolites, analyst B uses FDR<0.05 and reports 12, and analyst C adds Fold Change>1.5 and ends up with only 5. Which analyst is right? In practice, all three may be right — because they are answering different questions and managing different types of risk. A VIP-based list is optimized for class separation in a supervised model. An FDR-based list is optimized for controlling false discoveries across the full candidate set. A combined FC-and-FDR list is designed to retain features that are both biologically meaningful and statistically credible.
That is why, in differential feature screening omics, the real goal is not to generate the longest list. It is to generate the most defensible list for the next stage of validation. This is especially important in untargeted metabolomics studies, where different filtering orders can materially change the final candidates. Recent LC-MS work has shown that applying univariate filtering before or after PLS-DA can alter the selected features because noisy variables may distort the model and inflate VIP values [1]. Candidate screening, therefore, is not a contest for the smallest nominal p-value. It is a structured prioritization process that balances biological relevance, statistical confidence, and downstream validation cost.
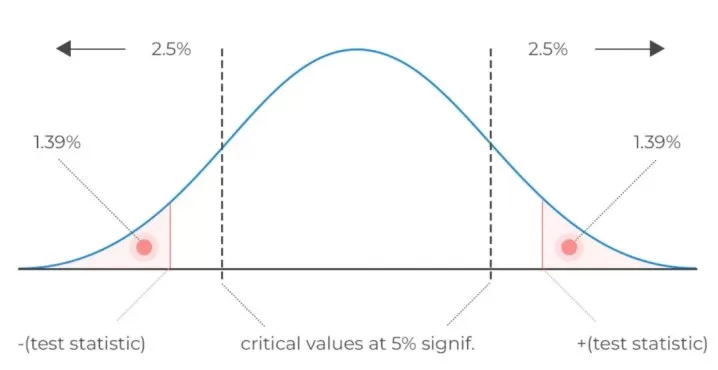
Figure 1. Overview of key statistical metrics — Fold Change, p-value, FDR, and VIP — used in differential feature screening for omics studies.
2. WHAT FOLD CHANGE, P-VALUE, FDR, AND VIP ACTUALLY TELL YOU
These four metrics are often used together, but they do not answer the same question. Fold Change measures effect size — it tells you whether the difference is large enough to matter biologically. The p-value evaluates whether the observed difference is unlikely to be due to random variation in a single comparison. FDR moves to the list level and asks whether the reported discoveries remain trustworthy after correcting for multiple testing. VIP, by contrast, reflects how strongly a feature contributes to group separation in a supervised multivariate model such as PLS-DA.
This is why the fold change vs p value omics debate is misleading. Fold Change and p-value are not competing metrics, and neither can replace FDR or VIP. They operate at different levels: effect magnitude, single-feature evidence, multiple-testing control, and multivariate contribution. In addition, the widely used VIP>1 threshold should be treated as a practical rule of thumb, not as definitive evidence that a molecule is a biomarker [2].
| Metric | Main question answered | Strength | Limitation if used alone |
|---|---|---|---|
| Fold Change | Is the difference large enough to be biologically meaningful? | Filters out trivial changes | Ignores variability and sample size |
| p-value | Is the difference unlikely to arise by chance in one test? | Detects statistical evidence | Inflates false positives in high-dimensional data |
| FDR | Can the full candidate list be trusted after multiple testing correction? | Controls false discovery burden | May exclude moderate but real signals in small cohorts |
| VIP | Does the feature contribute strongly to class separation in the model? | Captures covariance-driven discriminatory variables | Can be inflated by noisy features or overfitted models |
3. HOW THESE METHODS CAN BE COMBINED IN OMICS ANALYSIS
In practice, omics feature screening works best as a layered decision framework rather than a single cutoff. Different combinations are appropriate for different study goals. In early discovery, researchers often start with Fold Change and nominal p-value to capture a broad candidate space, especially when the goal is to identify pathways or generate hypotheses. In a more controlled shortlist stage, Fold Change is usually combined with FDR to keep only those features that are both biologically meaningful and statistically robust. When the goal is classification or subtype discrimination, analysts may further integrate VIP from a validated PLS-DA model to prioritize features that also contribute to multivariate separation.
The key principle is that each added metric should answer a missing question rather than repeat the same one. Fold Change addresses size, p-value addresses local uncertainty, FDR addresses project-level false-positive risk, and VIP addresses model-level contribution. Used together, they create a stronger evidence chain for omics differential analysis.
| Combination strategy | What it captures | Best used for | Main caution |
|---|---|---|---|
| FC + p-value | Large and individually significant changes | Early exploration, pathway discovery, pilot studies | Too many false positives if feature count is high |
| FC + FDR | Biologically meaningful and multiple-testing-corrected signals | Core shortlist generation in metabolomics or proteomics | May miss subtle but coordinated biology in underpowered datasets |
| FDR + VIP | Statistically credible features with multivariate discriminatory value | Biomarker panel discovery, response stratification, subtype classification | VIP must come from a rigorously validated model |
| FC + FDR + VIP | Integrated evidence across biology, statistics, and model contribution | Priority ranking for downstream validation and translational studies | Can become overly restrictive if thresholds are set too aggressively |
| Region-specific FC/FDR + spatial context | Differential signals with spatial interpretability | Spatial omics differential analysis, tumor microenvironment studies | Results depend heavily on region definition and comparison design |
4. WHY THIS MATTERS MORE IN OMICS THAN IN ORDINARY STATISTICS
Omics datasets are especially difficult because they combine high dimensionality with unstable inference conditions. Feature counts are large — often hundreds to thousands of metabolites and many thousands of proteins — so nominal significance thresholds can easily inflate false positives if multiple testing is not controlled. At the same time, sample sizes are often limited, making p-values more sensitive to variance structure, missing values, and outliers.
In proteomics, the challenge is even greater because missingness and technical variability are common, and batch correction is not just a preprocessing detail; it directly affects whether downstream significance calls are interpretable. Published studies on batch effects have shown that technical artifacts can become deeply misleading when they align with biological groups [3]. In both metabolomics and proteomics, univariate significance may also disagree with multivariate importance because covariance carries biological information that one-feature-at-a-time testing cannot capture.
The problem becomes even more complex in spatial assays. Once tissue architecture is considered, the relevant signal may exist in a specific region rather than in the whole-tissue average. This is why the best omics candidates are rarely the features with the most attractive single statistic. The strongest candidates are the ones that remain compelling across multiple layers of evidence.
5. METABOLOMICS EXAMPLE: TUMOR VERSUS MATCHED NORMAL TISSUE
A representative example comes from the Cancer Cell study that constructed an integrated metabolic atlas of clear cell renal cell carcinoma using 138 matched tumor and normal tissue pairs [4]. The study identified broad alterations in central carbon metabolism, one-carbon metabolism, and antioxidant pathways. It also showed that tumor progression and metastasis were associated with higher levels of glutathione and metabolites from the cysteine/methionine pathway.
This study is highly informative for metabolomics screening because it illustrates a common problem: pathway-level biology may be clear, while the value of any single metabolite as a validation target remains uneven. A metabolite may show a large Fold Change but still be a weak candidate if its within-group variation is high or if it fails to remain significant after multiple-testing correction. By contrast, a metabolite with a more moderate effect may deserve higher priority if it is consistent across matched pairs, passes FDR control, and supports a coherent biological mechanism.
That distinction matters in practice. Validation budgets are always limited. The most useful shortlist is therefore not the one with the most dramatic volcano plot, but the one that balances effect size, robustness, correction for multiple testing, and biological interpretability.
6. PROTEOMICS EXAMPLE: RESPONDERS VERSUS NON-RESPONDERS
The same logic applies even more clearly in proteomics biomarker screening, especially in response-stratified multi-omics studies. In such datasets, the biology is often driven by coordinated molecular programs rather than one dominant feature. Several proteins and metabolites may move together as part of the same response axis, and that coordinated behavior can be more informative than the smallest univariate p-value alone.
This is exactly why VIP can be useful: it highlights variables that contribute to multivariate class separation. However, that usefulness depends entirely on model quality. PLS-DA can overfit very easily, especially in small omics cohorts. As a result, VIP should only be used for prioritization after the model has been rigorously cross-validated and, ideally, stress-tested with permutation analysis. In other words, a feature with only moderate univariate significance may still be worth attention if it contributes strongly to a stable and well-validated multivariate model. This is one reason why panel-level thinking is often more informative than single-marker ranking in translational proteomics.
7. SPATIAL MULTI-OMICS EXAMPLE: WHY LOCATION CHANGES THE CANDIDATE
Spatial assays change the analytical question completely. Instead of asking whether a molecule differs on average, they ask where that difference occurs and in what tissue context. A representative example is the 2023 Nature Communications study that integrated spatial metabolomics, spatial lipidomics, and spatial transcriptomics in gastric cancer [5]. The authors identified a distinct immune-cell-dominated tumor-normal interface region with marked transcriptional and immunometabolic alterations, while also documenting substantial intratumoral heterogeneity.
For spatial omics differential analysis, this is not a minor technical refinement. It changes the unit of inference. A metabolite or protein that appears unremarkable in the tissue-wide average may become highly informative once the analysis is stratified into tumor core, invasive front, immune-rich interface, or other microregions. Spatial analysis therefore provides more than visual context. It improves mechanistic interpretability by showing whether a candidate is globally shifted, locally enriched, or restricted to a biologically meaningful niche.
Once location is added to the evidence chain, candidate ranking often changes substantially. In many cases, it also becomes more biologically convincing.
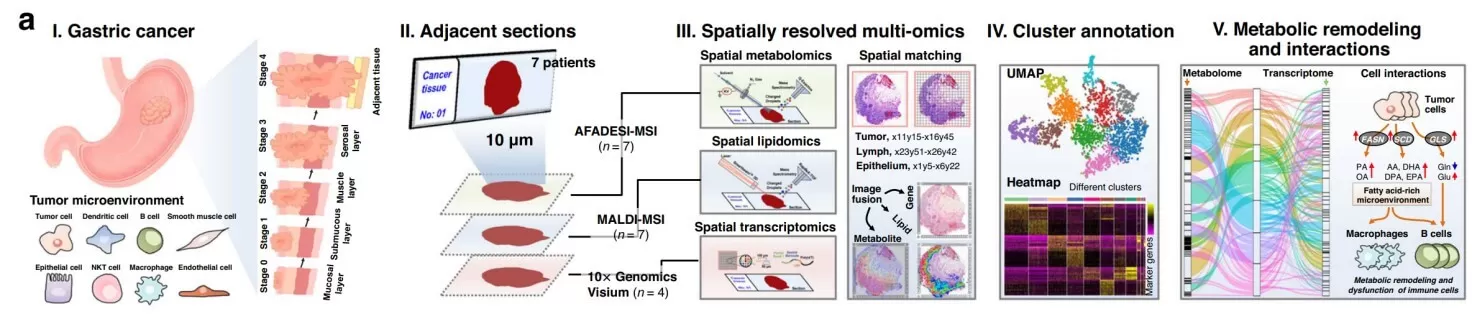
Figure 2. Integrated spatially resolved multi-omics for highlighting tumor metabolic remodeling. Image reproduced from Sun et al., 2023, Nature Communications, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
8. WHAT THE READER SHOULD TAKE AWAY
Any omics report built on a single metric should be treated cautiously. Raw p-values are risky because they ignore multiple testing. Fold Change alone is incomplete because a large shift may still be unstable. VIP>1 is not evidence of a biomarker by itself, because VIP reflects contribution within a supervised model and can be inflated if the model is not rigorously validated.
The most valuable candidates are those that pass several sequential questions. Is the effect large enough to matter biologically? Is it unlikely to be driven by random noise? Does it remain credible after multiple-testing correction? Does it contribute to stable multivariate separation rather than accidental classification? Once these questions are made explicit, feature screening becomes easier to justify in papers, grant applications, and industry-facing reports.
That is the real value of disciplined differential feature screening omics: it turns statistical output into a shortlist that is scientifically defensible and practically worth validating.
9. HOW WE SHOULD ACTUALLY DO IT
In omics, the best candidate is rarely the feature with the smallest raw p-value. The strongest candidates combine biological effect size, statistical robustness, multiple-testing control, and multivariate relevance. A strong screening strategy starts well before the first statistical test. Preprocessing and normalization directly affect Fold Change and p-values because they reshape the signal distribution and background variation. Batch-effect correction influences FDR stability because technical artifacts can otherwise be mistaken for biology. In proteomics, missing-value handling is especially important, as different imputation or harmonization strategies can change which proteins pass significance filtering.
For supervised analysis, VIP is useful only when the model itself is reliable. That means the number of latent variables must be justified, and the model must be evaluated by cross-validation and, ideally, permutation testing. In spatial studies, region definition is equally critical, because different segmentation strategies generate different biological contrasts and therefore different candidate lists.
The most professional omics workflow therefore builds a full evidence chain: sound study design, robust data acquisition, quality control, normalization, batch handling, differential analysis, integrated prioritization, and orthogonal validation. That is how feature screening moves from a statistical exercise to a decision framework that can support real biological interpretation and downstream investment.
Turn Omics Data Into More Confident Candidates
Differential feature screening is most useful when statistical evidence and biological relevance are evaluated together. That requires not only the right metrics, but also reliable preprocessing, quality control, and downstream interpretation.
MetwareBio supports metabolomics, proteomics, spatial omics, and multi-omics analysis to help researchers generate stronger candidate lists for validation.
Looking for support with your omics project? Contact us today.
Contact UsReferences
- Xu S, Bai C, Chen Y, et al. Comparing univariate filtration preceding and succeeding PLS-DA analysis on the differential variables/metabolites identified from untargeted LC-MS metabolomics data. Analytica Chimica Acta. 2024;1287:342103.
- Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society: Series B. 1995;57(1):289–300.
- Leek JT, et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nature Reviews Genetics. 2010;11:733–739.
- Hakimi AA, et al. An Integrated Metabolic Atlas of Clear Cell Renal Cell Carcinoma. Cancer Cell. 2016;29(1):104–116.
- Sun C, Wang A, Zhou Y, et al. Spatially resolved multi-omics highlights cell-specific metabolic remodeling and interactions in gastric cancer. Nature Communications. 2023;14:2692.


