Pearson vs Spearman Correlation: How to Choose the Right Method for Multi-Omics Data Analysis
Correlation analysis is one of the foundational tools in multi-omics data analysis, enabling researchers to uncover associations across genes, proteins, metabolites, and other molecular entities. However, its misuse remains prevalent, particularly in datasets with non-normal distributions, extreme values, or small sample sizes. Many studies default to Pearson correlation without considering these constraints, potentially producing misleading results, including false positives or false negatives. This article provides a systematic comparison of Pearson and Spearman correlation, emphasizing their theoretical distinctions, assumptions, and practical applications in genomics, proteomics, metabolomics, and multi-omics integration. By highlighting statistical pitfalls and guiding method selection, the discussion aims to equip researchers and bioinformaticians with best practices for reliable omics correlation analysis, improving the rigor and reproducibility of their studies.
1. Why Correlation Analysis Matters in Omics Data Analysis
Correlation measures the strength and direction of association between two variables, serving as a cornerstone for omics data analysis. In genomics, correlation networks reveal co-expression patterns that suggest shared regulatory mechanisms (Jiang et al., 2022); in proteomics and metabolomics, correlated abundance changes can indicate pathway coordination or biochemical interactions. The integration of multiple omics layers—multi-omics integration—relies heavily on correlation measures to bridge different molecular hierarchies and uncover systems-level biology.
However, high-dimensional omics datasets present unique challenges: non-normal distributions (common in RNA-seq count data), extreme outliers (from technical artifacts or biological variation), and limited sample sizes (particularly in clinical studies) all complicate correlation analysis. These challenges necessitate careful consideration of whether Pearson correlation—with its assumptions of linearity and normality—or the rank-based Spearman correlation is more appropriate for a given biological question (Rosa et al., 2022).
2. Pearson Correlation: Linear Association in Omics Data
Pearson correlation, introduced by Karl Pearson in the late 19th century, remains the most widely used measure of association in biological research. Its mathematical simplicity and interpretability make it an attractive choice, but its underlying assumptions require careful evaluation before application to omics data.
- Definition and Concept: The Pearson product-moment correlation coefficient (r) measures the strength and direction of a linear relationship between two continuous variables. Values range from -1 (perfect negative linear relationship) through 0 (no linear relationship) to +1 (perfect positive linear relationship).
- Mathematical Principle: The coefficient is calculated as the covariance of the two variables divided by the product of their standard deviations:
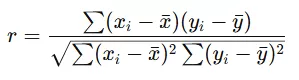
This formula essentially quantifies how far data points deviate from their means in a coordinated manner.
- Core Assumptions: Pearson correlation requires that: 1) Data approximately follow a bivariate normal distribution (at least unimodal and symmetric); 2) The relationship between variables is linear; 3) No substantial outliers are present (outliers can dramatically inflate or deflate correlation estimates)
- Applications in Multi-Omics: Pearson correlation is appropriate when relationships are known to be linear, such as in technical replicate comparisons for quality assessment. After appropriate normalization and variance-stabilizing transformation of expression data, Pearson correlation can effectively capture linear relationships.
3. Spearman Correlation: Rank-Based Analysis for Multi-Omics
Spearman's rank correlation offers a robust alternative that frees analysts from the distributional assumptions required by Pearson. By operating on data ranks rather than raw values, it provides reliable inference across a wider range of data scenarios common in omics research.
- Definition and Concept: Spearman's rank correlation coefficient (ρ or rₛ) measures the strength and direction of a monotonic relationship between two variables—one that consistently increases or decreases, though not necessarily at a constant rate. Like Pearson, it ranges from -1 to +1.
- Mathematical Principle: Raw data values are converted to ranks (the smallest value receives rank 1, the next rank 2, etc.), and Pearson correlation is calculated on these ranks. An equivalent formula based on rank differences is:
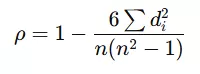
where didi represents the difference between the ranks of each observation pair.
- Core Advantages: 1) Makes no distributional assumptions (nonparametric); 2) Resistant to outliers—extreme values lose their disproportionate influence after rank transformation 3) Captures monotonic nonlinear relationships (e.g., exponential growth, saturation kinetics)
- Applications in Multi-Omics: Spearman correlation excels in common omics scenarios. Gene expression data, particularly from RNA-seq, typically exhibit skewed distributions, making Spearman the preferred choice for constructing co-expression networks. In metabolomics, where concentration data often contain extreme values, Spearman provides more stable estimates. For exploratory analyses where relationship shapes are unknown, Spearman offers a conservative starting point.
4. Pearson vs Spearman: Key Differences and Selection Guidelines
The choice between Pearson and Spearman correlation fundamentally depends on data characteristics and research questions. Neither method is universally superior; each has domains where it performs optimally. The following comparison highlights key distinctions to guide method selection.
|
Feature |
Pearson Correlation |
Spearman Correlation |
|
Relationship Type |
Linear relationships |
Monotonic relationships (linear or nonlinear) |
|
Data Type |
Continuous variables, approximate normality |
Continuous or ordinal variables, no distributional requirements |
|
Parameter/Nonparametric |
Parametric method |
Nonparametric method |
|
Outlier Sensitivity |
Highly sensitive |
Robust |
|
Statistical Power |
Higher power for normal, linear data |
Higher power for heavy-tailed or nonlinear data |
|
Variability |
Lower variability in normal, light-tailed data |
Lower variability in non-normal data (up to 20% reduction) |
How to Choose Based on Data Characteristics:
1) Visualize relationships with scatter plots to assess linearity and identify potential outliers.
2) Examine data distributions (using Shapiro-Wilk tests or Q-Q plots) and check for extreme values.
3) Apply the following decision rules:
- If relationships appear linear, outliers are absent, and data approximate normality, choose Pearson correlation
- If relationships are monotonic but nonlinear, outliers are present, or distributions are heavily skewed, choose Spearman correlation
- If complex non-monotonic patterns (e.g., U-shaped) are suspected, consider advanced methods like distance correlation
4) Interpreting Disagreement Between Methods:
- High Pearson, low Spearman: Suggests a linear relationship driven by outliers—the Pearson estimate may be artificially inflated.
- High Spearman, low Pearson: Indicates a monotonic but nonlinear relationship that Pearson fails to capture.
5) Advanced Extensions: For complex nonlinear patterns, distance correlation offers a distribution-free measure capable of detecting any type of dependence, not just monotonic relationships (Hou et al., 2022). Maximal information coefficient (MIC) provides another alternative for capturing a wide range of association patterns (de Reshef et al., 2012; Winter et al., 2016).
5. Practical Applications: Choosing Pearson or Spearman in Multi-Omics
The choice between Pearson and Spearman correlation has direct implications for multi-omics data analysis. Each method’s assumptions and strengths dictate its suitability for different omics types, such as transcriptomics, proteomics, metabolomics, or microbiome data. Understanding these distinctions allows analysts to identify biologically meaningful associations while minimizing false positives or biases caused by inappropriate correlation measures.
Genomics and Transcriptomics Correlation Analysis
Transcriptomic and genomics datasets, including bulk RNA‑seq and single‑cell RNA‑seq, are dominated by count data with skewed distributions, extensive heterogeneity, and technical noise such as dropouts and low counts. These properties frequently violate the normality and linearity assumptions of Pearson correlation, resulting in biased or unstable estimates when used naively in co‑expression and regulatory network analysis. Non‑parametric Spearman correlation, by ranking expression values, provides a robust alternative that is insensitive to distributional shape and outliers, making it more suited to capture monotonic relationships in high‑dimensional transcriptomic data. For example, in a recent large‑scale single‑cell co‑expression benchmarking study, Spearman correlation of log normalized RNA‑seq expression was explicitly compared against other metrics as part of evaluating co‑expression inference methods, demonstrating its utility in identifying monotonic gene–gene associations in noisy transcriptomic measurements (Su et al., 2023).
Proteomics and Metabolomics Correlation Analysis
Proteomics and metabolomics datasets are characterized by wide dynamic ranges, skewed concentration distributions, and often non‑Gaussian features resulting from biological regulation and measurement processes. In many multi‑omics contexts, the relationship between protein abundance or metabolite levels and other omics layers may be non‑linear or influenced by outliers, which challenges strictly linear correlation metrics. A pragmatic strategy is to choose Pearson correlation when data are approximately normalized and linear relationships are expected (e.g., technical replicates or replicated conditions), but prioritize Spearman correlation when investigating broad co‑variation patterns across diverse samples or when distributions show heavy tails. In a recent open‑access multi‑omics study of human aortic smooth muscle cells exposed to chronic hyperglycemia, Spearman’s correlation coefficient was explicitly used to quantify the association between mRNA abundance from RNA‑seq and corresponding protein levels measured by mass spectrometry, revealing moderate but significant transcript–protein concordance across conditions (Bohara et al., 2024).
 between ECM proteins and mRNA levels across various gl_1774008551_WNo_685d528.webp)
Correlation (Spearman correlation coefficient) between ECM proteins and mRNA levels across various glucose conditions and T2DM patient samples for 448 genes.
Image reproduced from Bohara et al., 2024, Journal of Biological Engineering
Multi‑Omics Integration Analysis (Microbiome + Metabolome)
Multi‑omics integration, such as linking microbiome composition with host metabolome profiles, further complicates correlation analysis due to compositional constraints, high sparsity, and distinct measurement scales across datasets. Pearson correlation may be inappropriate when relative abundance and zero‑inflation dominate, whereas Spearman’s rank correlation is frequently employed to capture monotonic associations between microbial taxa and metabolite levels without assuming linearity or normality. In a recent open‑access hepatocellular carcinoma (HCC) study integrating gut microbiota and serum metabolite profiles, Spearman’s rank correlation was systematically applied to compute taxa–metabolite association coefficients, enabling the identification of key microbial species linked to differential metabolites as potential biomarkers, while explicitly controlling for multiple testing (Li et al., 2023). This approach provided biologically interpretable inter‑omics associations in the noisy and high‑dimensional microbiome–metabolome space.
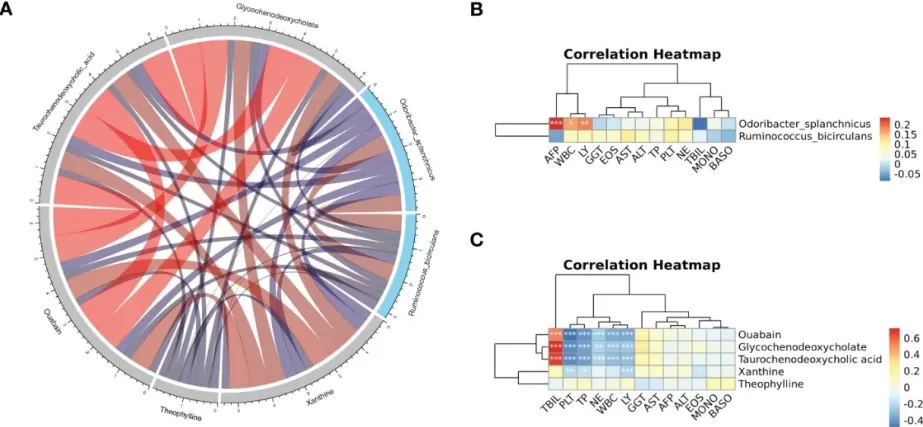
Integrated Correlation Analysis of Metabolites, Microbiota, and Clinical Traits
(A) Spearman correlation chord diagram of key species and key metabolites. (B, C) Pearson correlation cluster heatmap depicting the relationships between the key metabolites, the key species and the clinical indicators.
Image reproduced from Li et al., 2023, Frontiers in cellular and infection microbiology, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
6. Practical Tools: Implementing Pearson and Spearman Correlation
Implementing Pearson and Spearman correlation analysis in modern bioinformatics workflows requires familiarity with standard software tools and awareness of best practices for result interpretation.
R Implementation:
- Basic correlation: cor(x, y, method = "pearson") or cor(x, y, method = "spearman")
- With significance testing: cor.test(x, y, method = "pearson") provides both correlation coefficient and p-value
- For correlation matrices: cor(df, method = "spearman") calculates all pairwise correlations
Python Implementation:
- Using SciPy: scipy.stats.pearsonr(x, y) returns correlation coefficient and p-value
- Spearman equivalent: scipy.stats.spearmanr(x, y)
- For multiple comparisons: pandas.DataFrame.corr(method='spearman') computes correlation matrices
7. Common Misconceptions and Best Practices in Correlation Analysis
Correlation analysis in omics research is often misunderstood, leading to frequent misinterpretations. A common misconception is that non-normal or skewed data automatically necessitates Spearman correlation. While Spearman is robust to non-normality and outliers, Pearson correlation can still be appropriate when relationships are linear and outliers are controlled, offering higher statistical power. Another frequent error is assuming that high correlation coefficients directly indicate biological co-expression or functional interactions. In reality, correlations can arise from confounding factors such as batch effects, technical variability, or sample heterogeneity, potentially producing spurious associations. Additionally, it is incorrect to assume that Pearson and Spearman correlations will always yield similar results; monotonic non-linear relationships may produce high Spearman coefficients with near-zero Pearson values, highlighting the need to understand the underlying data distribution and relationship type.
Best practices in multi-omics correlation analysis involve careful data preprocessing, method selection, and validation. Data normalization, log-transformation, and outlier assessment help meet method assumptions and reduce bias. Visualization through scatterplots and heatmaps can guide the choice between Pearson and Spearman correlation based on observed linearity and monotonic trends. For high-dimensional datasets, multiple testing correction and integration with confounder-adjusted approaches, such as partial correlation or tools like CorrAdjust, improve reliability. When complex non-linear associations are suspected, advanced measures like distance correlation or maximal information coefficient provide complementary insights. Adopting these strategies ensures robust and reproducible correlation analysis across genomics, proteomics, metabolomics, and integrated multi-omics workflows, enhancing the interpretability and biological relevance of the findings.
Reference:
1. Jiang, Y. H., Long, J., Zhao, Z. B., Li, L., Lian, Z. X., Liang, Z., & Wu, J. R. (2022). Gene co-expression network based on part mutual information for gene-to-gene relationship and gene-cancer correlation analysis. BMC bioinformatics, 23(1), 194. https://doi.org/10.1186/s12859-022-04732-9
2. Rosa, J. C. D., Aleman, J. O., Mohabir, J., Liang, Y., Breslow, J. L., & Holt, P. R. (2022). The Application of Spearman Partial Correlation for Screening Predictors of Weight Loss in a Multiomics Dataset. Omics : a journal of integrative biology, 26(12), 660–670. https://doi.org/10.1089/omi.2022.0135
3. Hou, J., Ye, X., Feng, W., Zhang, Q., Han, Y., Liu, Y., Li, Y., & Wei, Y. (2022). Distance correlation application to gene co-expression network analysis. BMC bioinformatics, 23(1), 81. https://doi.org/10.1186/s12859-022-04609-x
4. Reshef, D. N., Reshef, Y. A., Finucane, H. K., Grossman, S. R., McVean, G., Turnbaugh, P. J., Lander, E. S., Mitzenmacher, M., & Sabeti, P. C. (2011). Detecting novel associations in large data sets. Science (New York, N.Y.), 334(6062), 1518–1524. https://doi.org/10.1126/science.1205438
5. de Winter, J. C., Gosling, S. D., & Potter, J. (2016). Comparing the Pearson and Spearman correlation coefficients across distributions and sample sizes: A tutorial using simulations and empirical data. Psychological methods, 21(3), 273–290. https://doi.org/10.1037/met0000079
6. Su, C., Xu, Z., Shan, X., Cai, B., Zhao, H., & Zhang, J. (2023). Cell-type-specific co-expression inference from single cell RNA-sequencing data. Nature communications, 14(1), 4846. https://doi.org/10.1038/s41467-023-40503-7
7. Bohara, S., Bagheri, A., Ertugral, E. G., Radzikh, I., Sandlers, Y., Jiang, P., & Kothapalli, C. R. (2024). Integrative analysis of gene expression, protein abundance, and metabolomic profiling elucidates complex relationships in chronic hyperglycemia-induced changes in human aortic smooth muscle cells. Journal of biological engineering, 18(1), 61. https://doi.org/10.1186/s13036-024-00457-w
8. Li, X., Yi, Y., Wu, T., Chen, N., Gu, X., Xiang, L., Jiang, Z., Li, J., & Jin, H. (2023). Integrated microbiome and metabolome analysis reveals the interaction between intestinal flora and serum metabolites as potential biomarkers in hepatocellular carcinoma patients. Frontiers in cellular and infection microbiology, 13, 1170748. https://doi.org/10.3389/fcimb.2023.1170748
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


